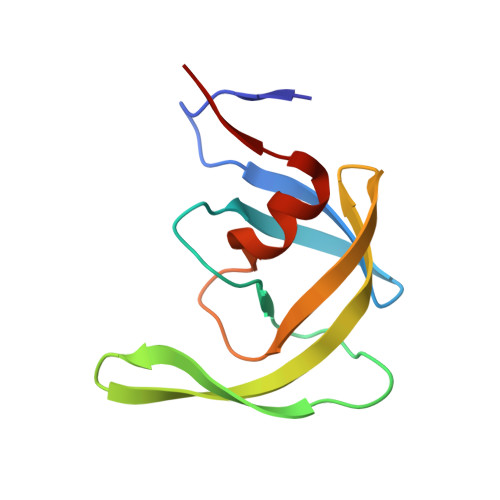
Insights into the mechanism of drug resistance: X-ray structure analysis of multi-drug resistant HIV-1 protease ritonavir complex.
Liu, Z., Yedidi, R.S., Wang, Y., Dewdney, T.G., Reiter, S.J., Brunzelle, J.S., Kovari, I.A., Kovari, L.C.(2013) Biochem Biophys Res Commun 431: 232-238
- PubMed: 23313846 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2012.12.127
- Primary Citation Related Structures:
4EYR - PubMed Abstract:
Ritonavir (RTV) is a first generation HIV-1 protease inhibitor with rapidly emerging drug resistance. Mutations at residues 46, 54, 82 and 84 render the HIV-1 protease drug resistant against RTV. We report the crystal structure of multi-drug resistant (MDR) 769 HIV-1 protease (carrying resistant mutations at residues 10, 36, 46, 54, 62, 63, 71, 82, 84 and 90) complexed with RTV and the in vitro enzymatic IC(50) of RTV against MDR HIV-1 protease. The structural and functional studies demonstrate significant drug resistance of MDR HIV-1 protease against RTV, arising from reduced hydrogen bonds and Van der Waals interactions between RTV and MDR HIV-1 protease.
- Department of Biochemistry and Molecular Biology, School of Medicine, Wayne State University, Detroit, MI 48201, USA.
Organizational Affiliation: