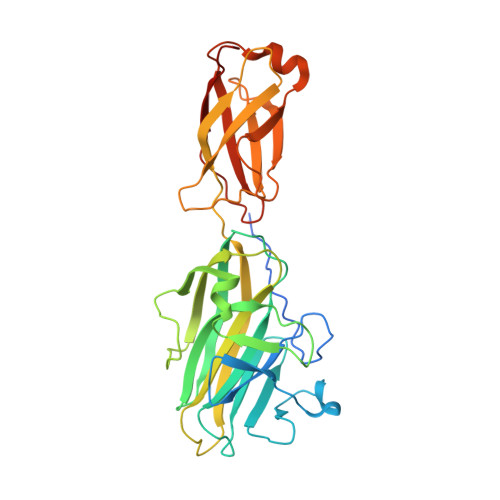
The Extracellular Protein Factor Epf from Streptococcus pyogenes Is a Cell Surface Adhesin That Binds to Cells through an N-terminal Domain Containing a Carbohydrate-binding Module.
Linke, C., Siemens, N., Oehmcke, S., Radjainia, M., Law, R.H., Whisstock, J.C., Baker, E.N., Kreikemeyer, B.(2012) J Biological Chem 287: 38178-38189
- PubMed: 22977243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.376434
- Primary Citation Related Structures:
4ES8, 4ES9 - PubMed Abstract:
Streptococcus pyogenes is an exclusively human pathogen. Streptococcal attachment to and entry into epithelial cells is a prerequisite for a successful infection of the human host and requires adhesins. Here, we demonstrate that the multidomain protein Epf from S. pyogenes serotype M49 is a streptococcal adhesin. An epf-deficient mutant showed significantly decreased adhesion to and internalization into human keratinocytes. Cell adhesion is mediated by the N-terminal domain of Epf (EpfN) and increased by the human plasma protein plasminogen. The crystal structure of EpfN, solved at 1.6 Å resolution, shows that it consists of two subdomains: a carbohydrate-binding module and a fibronectin type III domain. Both fold types commonly participate in ligand receptor and protein-protein interactions. EpfN is followed by 18 repeats of a domain classified as DUF1542 (domain of unknown function 1542) and a C-terminal cell wall sorting signal. The DUF1542 repeats are not involved in adhesion, but biophysical studies show they are predominantly α-helical and form a fiber-like stalk of tandem DUF1542 domains. Epf thus conforms with the widespread family of adhesins known as MSCRAMMs (microbial surface components recognizing adhesive matrix molecules), in which a cell wall-attached stalk enables long range interactions via its adhesive N-terminal domain.
- Maurice Wilkins Centre for Molecular Biodiscovery and School of Biological Sciences, University of Auckland, Auckland 1142, New Zealand.
Organizational Affiliation: