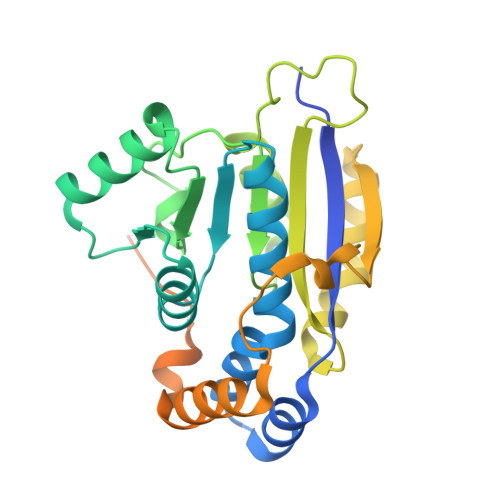
Structure of the C-terminal half of human XPB helicase and the impact of the disease-causing mutation XP11BE.
Hilario, E., Li, Y., Nobumori, Y., Liu, X., Fan, L.(2013) Acta Crystallogr D Biol Crystallogr 69: 237-246
- PubMed: 23385459 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912045040
- Primary Citation Related Structures:
4ERN - PubMed Abstract:
XPB is a DNA-dependent helicase and a subunit of the TFIIH complex required for both transcription and DNA repair. XPB contains four domains: an N-terminal domain, two conserved helicase domains (HD1 and HD2) and a C-terminal extension. The C-terminal extension is important for DNA repair since the phosphorylation of Ser751 inhibits 5'-incision by ERCC1-XPF endonuclease. A disease-causing frameshift mutation (XP11BE) that changes the last 42 amino acids of XPB causes manifestations including impaired DNA repair and deficient transcription. Here, the crystal structure of the C-terminal half of XPB (residues 494-782) is reported at 1.8 Å resolution. The structure contained the conserved XPB HD2 and a C-terminal extension which shares structural similarity with RIG-I, leading to a structural model of the XPF-XPB-DNA complex for 5' incision during DNA repair. A mutation mimicking the XP11BE mutation produced the much less soluble mutant XPBm(494-781). Western blotting results confirmed that the intracellular levels of XPB and other TFIIH subunits in XP11BE patient cells were much lower than those from the healthy parents. Together, these results indicate that the XP11BE mutation not only divests the XPF-interaction motif, impairing DNA repair, but also reduces XPB solubility, leading to a lower intracellular level of TFIIH and deficient transcription.
- Department of Biochemistry, University of California, Riverside, Riverside, CA 92521, USA.
Organizational Affiliation: