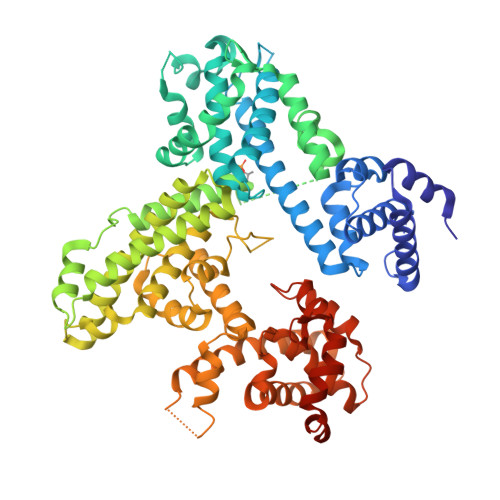
Structures of inactive retinoblastoma protein reveal multiple mechanisms for cell cycle control.
Burke, J.R., Hura, G.L., Rubin, S.M.(2012) Genes Dev 26: 1156-1166
- PubMed: 22569856 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.189837.112
- Primary Citation Related Structures:
4ELJ, 4ELL - PubMed Abstract:
Cyclin-dependent kinase (Cdk) phosphorylation of the Retinoblastoma protein (Rb) drives cell proliferation through inhibition of Rb complexes with E2F transcription factors and other regulatory proteins. We present the first structures of phosphorylated Rb that reveal the mechanism of its inactivation. S608 phosphorylation orders a flexible "pocket" domain loop such that it mimics and directly blocks E2F transactivation domain (E2F(TD)) binding. T373 phosphorylation induces a global conformational change that associates the pocket and N-terminal domains (RbN). This first multidomain Rb structure demonstrates a novel role for RbN in allosterically inhibiting the E2F(TD)-pocket association and protein binding to the pocket "LxCxE" site. Together, these structures detail the regulatory mechanism for a canonical growth-repressive complex and provide a novel example of how multisite Cdk phosphorylation induces diverse structural changes to influence cell cycle signaling.
- Department of Chemistry and Biochemistry, University of California at Santa Cruz, Santa Cruz, California 95064, USA.
Organizational Affiliation: