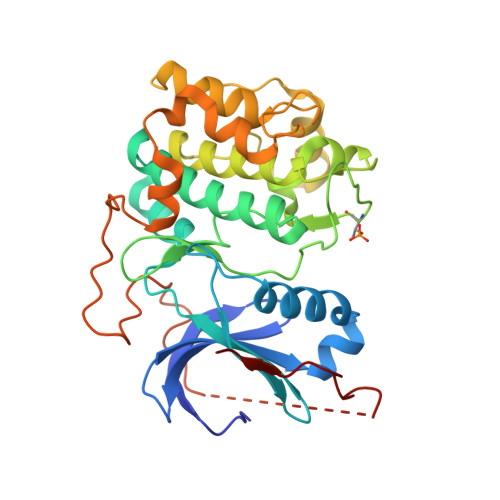
An ATP-Site On-Off Switch That Restricts Phosphatase Accessibility of Akt.
Lin, K., Lin, J., Wu, W.I., Ballard, J., Lee, B.B., Gloor, S.L., Vigers, G.P., Morales, T.H., Friedman, L.S., Skelton, N., Brandhuber, B.J.(2012) Sci Signal 5: ra37-ra37
- PubMed: 22569334 Search on PubMed
- DOI: https://doi.org/10.1126/scisignal.2002618
- Primary Citation Related Structures:
4EKK, 4EKL - PubMed Abstract:
The protein serine-threonine kinase Akt undergoes a substantial conformational change upon activation, which is induced by the phosphorylation of two critical regulatory residues, threonine 308 and serine 473. Paradoxically, treating cells with adenosine 5'-triphosphate (ATP)-competitive inhibitors of Akt results in increased phosphorylation of both residues. We show that binding of ATP-competitive inhibitors stabilized a conformation in which both phosphorylated sites were inaccessible to phosphatases. ATP binding also produced this protection of the phosphorylated sites, whereas interaction with its hydrolysis product adenosine 5'-diphosphate (ADP) or allosteric Akt inhibitors resulted in increased accessibility of these phosphorylated residues. ATP-competitive inhibitors mimicked ATP by targeting active Akt. Forms of Akt activated by an oncogenic mutation or myristoylation were more potently inhibited by the ATP-competitive inhibitors than was wild-type Akt. These data support a new model of kinase regulation, wherein nucleotides modulate an on-off switch in Akt through conformational changes, which is disrupted by ATP-competitive inhibitors.
- Genentech Inc., South San Francisco, CA 94080, USA. klin@gene.com
Organizational Affiliation: