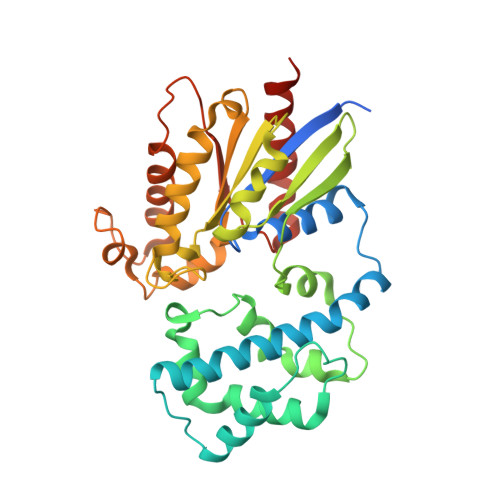
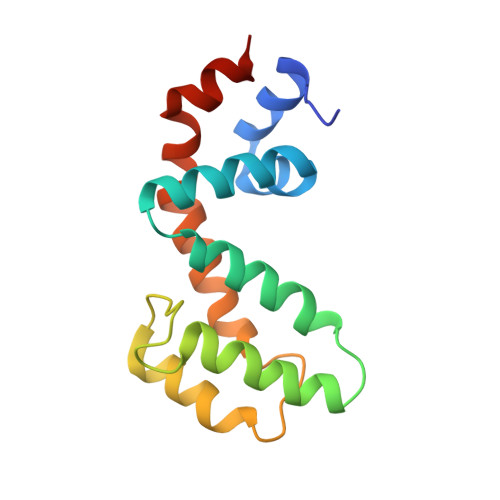
Structural and functional analysis of the regulator of G protein signaling 2-g alpha q complex.
Nance, M.R., Kreutz, B., Tesmer, V.M., Sterne-Marr, R., Kozasa, T., Tesmer, J.J.(2013) Structure 21: 438-448
- PubMed: 23434405 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.12.016
- Primary Citation Related Structures:
4EKC, 4EKD - PubMed Abstract:
The heterotrimeric G protein Gαq is a key regulator of blood pressure, and excess Gαq signaling leads to hypertension. A specific inhibitor of Gαq is the GTPase activating protein (GAP) known as regulator of G protein signaling 2 (RGS2). The molecular basis for how Gαq/11 subunits serve as substrates for RGS proteins and how RGS2 mandates its selectivity for Gαq is poorly understood. In crystal structures of the RGS2-Gαq complex, RGS2 docks to Gαq in a different orientation from that observed in RGS-Gαi/o complexes. Despite its unique pose, RGS2 maintains canonical interactions with the switch regions of Gαq in part because its α6 helix adopts a distinct conformation. We show that RGS2 forms extensive interactions with the α-helical domain of Gαq that contribute to binding affinity and GAP potency. RGS subfamilies that do not serve as GAPs for Gαq are unlikely to form analogous stabilizing interactions.
- Life Sciences Institute, University of Michigan, Ann Arbor, MI 48109-2216, USA.
Organizational Affiliation: