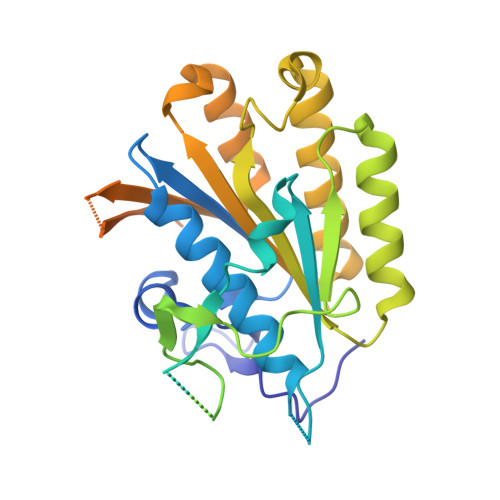
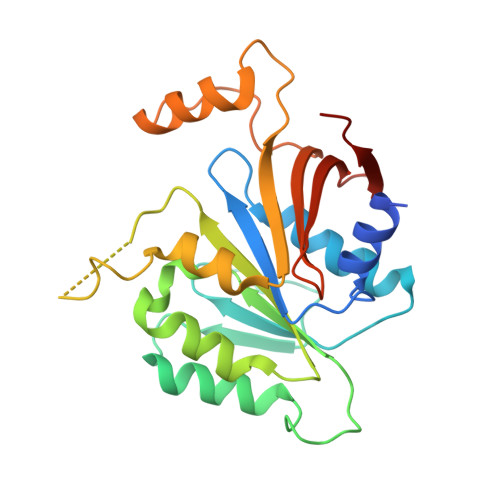
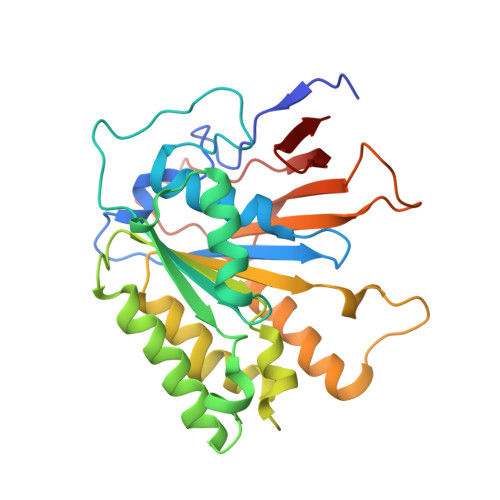
Crystal structure of elongator subcomplex Elp4-6
Lin, Z., Zhao, W., Diao, W., Xie, X., Wang, Z., Zhang, J., Shen, Y., Long, J.(2012) J Biological Chem 287: 21501-21508
- PubMed: 22556426
- DOI: https://doi.org/10.1074/jbc.M112.341560
- Primary Citation Related Structures:
4EJS - PubMed Abstract:
Elongator is a multiprotein complex composed of two subcomplexes, Elp1-3 and Elp4-6. Elongator is highly conserved between yeast and humans and plays an important role in RNA polymerase II-mediated transcriptional elongation and many other processes, including cytoskeleton organization, exocytosis, and tRNA modification. Here, we determined the crystal structure of the Elp4-6 subcomplex of yeast. The overall structure of Elp4-6 revealed that Elp6 acts as a bridge to assemble Elp4 and Elp5. Detailed structural and sequence analyses revealed that each subunit in the Elp4-6 subcomplex forms a RecA-ATPase-like fold, although it lacks the key sequence signature of ATPases. Site-directed mutagenesis and biochemical analyses indicated that the Elp4-6 subcomplex can assemble into a hexameric ring-shaped structure in vitro and in vivo. Furthermore, GST pulldown assays showed that the ring-shaped assembly of the Elp4-6 subcomplex is important for its specific histone H3 binding. Our results may shed light on the substrate recognition and assembly of the holo-Elongator complex.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, 94 Weijin Road, Tianjin 300071, China.
Organizational Affiliation: