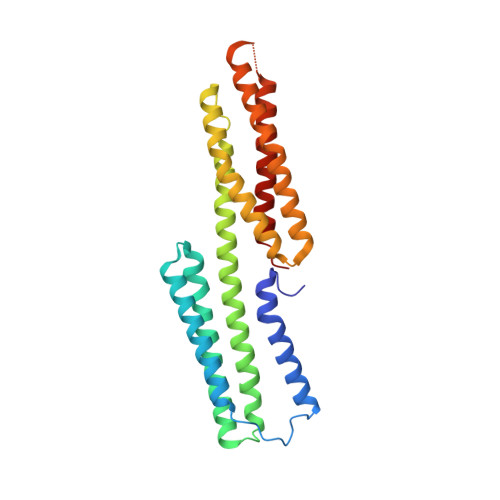
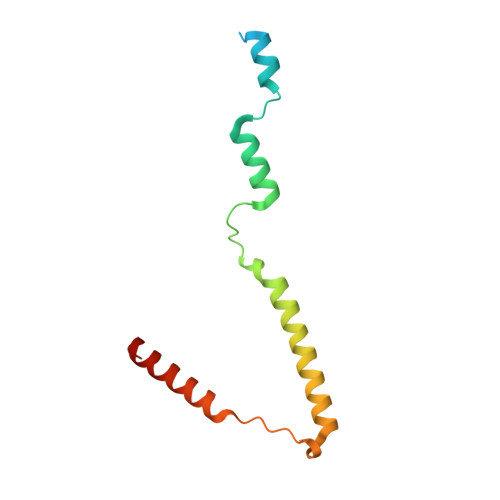
alpha-Catenin unfurls upon binding to vinculin
Rangarajan, E.S., Izard, T.(2012) J Biological Chem 287: 18492-18499
- PubMed: 22493458 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.351023
- Primary Citation Related Structures:
4EHP - PubMed Abstract:
Adherens junctions (AJs) are essential for cell-cell contacts, morphogenesis, and the development of all higher eukaryotes. AJs are formed by calcium-dependent homotypic interactions of the ectodomains of single membrane-pass cadherin family receptors. These homotypic interactions in turn promote binding of the intracellular cytoplasmic tail domains of cadherin receptors with β-catenin, a multifunctional protein that plays roles in both transcription and AJs. The cadherin receptor-β-catenin complex binds to the cytoskeletal protein α-catenin, which is essential for both the formation and the stabilization of these junctions. Precisely how α-catenin contributes to the formation and stabilization of AJs is hotly debated, although the latter is thought to involve its interactions with the cytoskeletal protein vinculin. Here we report the crystal structure of the vinculin binding domain (VBD) of α-catenin in complex with the vinculin head domain (Vh1). This structure reveals that α-catenin is in a unique unfurled mode allowing dimer formation when bound to vinculin. Finally, binding studies suggest that vinculin must be in an activated state to bind to α-catenin and that this interaction is stabilized by the formation of a ternary α-catenin-vinculin-F-actin complex, which can be formed via the F-actin binding domain of either protein. We propose a feed-forward model whereby α-catenin-vinculin interactions promote their binding to the actin cytoskeleton to stabilize AJs.
- Department of Cancer Biology, The Scripps Research Institute, Jupiter, Florida 33458, USA.
Organizational Affiliation: