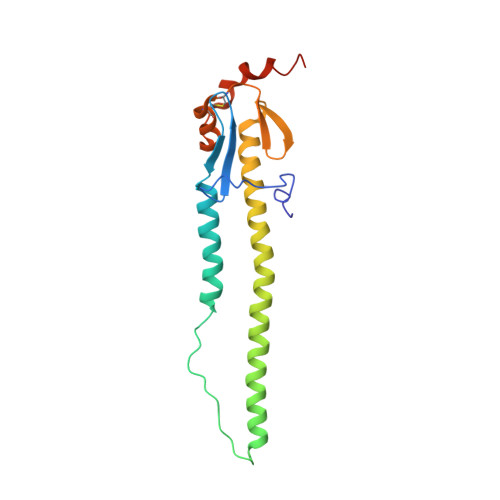
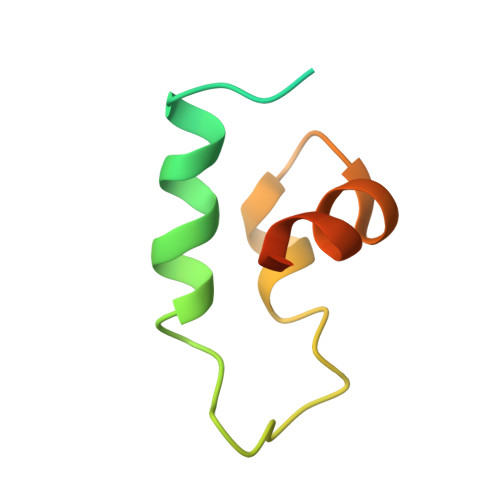
Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing.
Whitehead, T.A., Chevalier, A., Song, Y., Dreyfus, C., Fleishman, S.J., De Mattos, C., Myers, C.A., Kamisetty, H., Blair, P., Wilson, I.A., Baker, D.(2012) Nat Biotechnol 30: 543-548
- PubMed: 22634563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nbt.2214
- Primary Citation Related Structures:
4EEF - PubMed Abstract:
We show that comprehensive sequence-function maps obtained by deep sequencing can be used to reprogram interaction specificity and to leapfrog over bottlenecks in affinity maturation by combining many individually small contributions not detectable in conventional approaches. We use this approach to optimize two computationally designed inhibitors against H1N1 influenza hemagglutinin and, in both cases, obtain variants with subnanomolar binding affinity. The most potent of these, a 51-residue protein, is broadly cross-reactive against all influenza group 1 hemagglutinins, including human H2, and neutralizes H1N1 viruses with a potency that rivals that of several human monoclonal antibodies, demonstrating that computational design followed by comprehensive energy landscape mapping can generate proteins with potential therapeutic utility.
- Department of Biochemistry, University of Washington, Seattle, Washington, USA.
Organizational Affiliation: