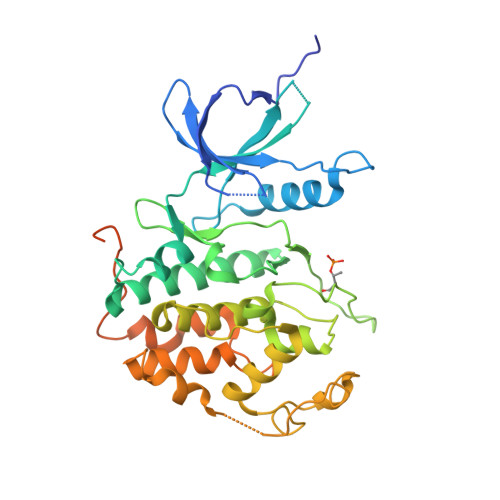
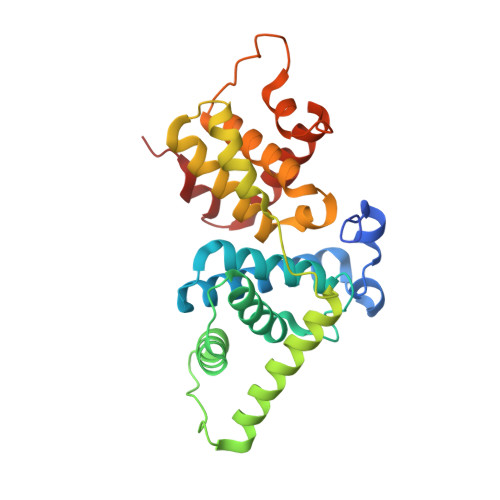
The CDK9 tail determines the reaction pathway of positive transcription elongation factor b.
Baumli, S., Hole, A.J., Wang, L.Z., Noble, M.E., Endicott, J.A.(2012) Structure 20: 1788-1795
- PubMed: 22959624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2012.08.011
- Primary Citation Related Structures:
4EC8, 4EC9 - PubMed Abstract:
CDK9, the kinase of positive transcription elongation factor b (P-TEFb), stimulates transcription elongation by phosphorylating RNA polymerase II and transcription elongation factors. Using kinetic analysis of a human P-TEFb complex consisting of CDK9 and cyclin T, we show that the CDK9 C-terminal tail sequence is important for the catalytic mechanism and imposes an ordered binding of substrates and release of products. Crystallographic analysis of a CDK9/cyclin T complex in which the C-terminal tail partially blocks the ATP binding site reveals a possible reaction intermediate. Biochemical characterization of CDK9 mutants supports a model in which the CDK9 tail cycles through different conformational states. We propose that this mechanism is critical for the pattern of CTD Ser2 phosphorylation on actively transcribed genes.
- Northern Institute for Cancer Research, Newcastle University, Newcastle upon Tyne NE2 4HH, UK. sbaumli@gmx.ch
Organizational Affiliation: