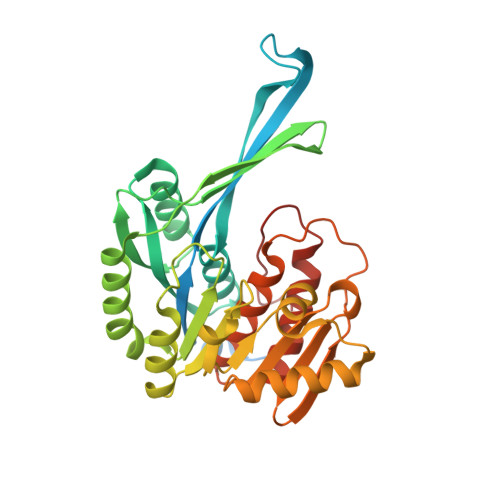
Structural-functional studies of Burkholderia cenocepacia D-glycero-beta-D-manno-heptose 7-phosphate kinase (HldA) and characterization of inhibitors with antibiotic adjuvant and antivirulence properties.
Lee, T.W., Verhey, T.B., Antiperovitch, P.A., Atamanyuk, D., Desroy, N., Oliveira, C., Denis, A., Gerusz, V., Drocourt, E., Loutet, S.A., Hamad, M.A., Stanetty, C., Andres, S.N., Sugiman-Marangos, S., Kosma, P., Valvano, M.A., Moreau, F., Junop, M.S.(2013) J Med Chem 56: 1405-1417
- PubMed: 23256532 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm301483h
- Primary Citation Related Structures:
4E84, 4E8W, 4E8Y, 4E8Z - PubMed Abstract:
As an essential constituent of the outer membrane of Gram-negative bacteria, lipopolysaccharide contributes significantly to virulence and antibiotic resistance. The lipopolysaccharide biosynthetic pathway therefore serves as a promising therapeutic target for antivirulence drugs and antibiotic adjuvants. Here we report the structural-functional studies of D-glycero-β-D-manno-heptose 7-phosphate kinase (HldA), an absolutely conserved enzyme in this pathway, from Burkholderia cenocepacia. HldA is structurally similar to members of the PfkB carbohydrate kinase family and appears to catalyze heptose phosphorylation via an in-line mechanism mediated mainly by a conserved aspartate, Asp270. Moreover, we report the structures of HldA in complex with two potent inhibitors in which both inhibitors adopt a folded conformation and occupy the nucleotide-binding sites. Together, these results provide important insight into the mechanism of HldA-catalyzed heptose phosphorylation and necessary information for further development of HldA inhibitors.
- Department of Biochemistry and Biomedical Sciences and Michael G. DeGroote Institute for Infectious Disease Research, McMaster University, 1280 Main Street West, Hamilton, Ontario, L8S 4K1, Canada.
Organizational Affiliation: