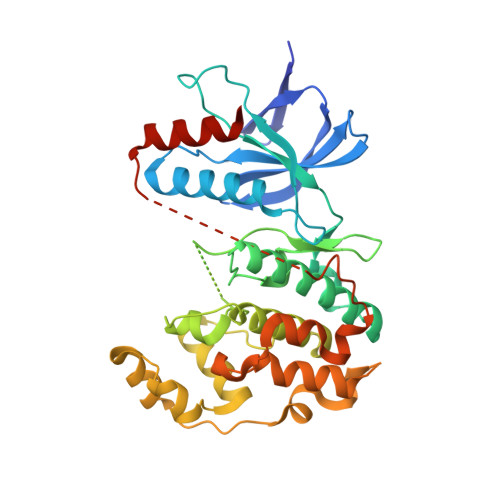
Identification of an Adamantyl Azaquinolone JNK Selective Inhibitor.
Haynes, N.E., Scott, N.R., Chen, L.C., Janson, C.A., Li, J.K., Lukacs, C.M., Railkar, A., Tozzo, E., Whittard, T., Brown, N.F., Cheung, A.W.(2012) ACS Med Chem Lett 3: 764-768
- PubMed: 24900545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml300175c
- Primary Citation Related Structures:
4E73 - PubMed Abstract:
3-[4-((1S,2S,3R,5S,7S)-5-Hydroxyadamantan-2-ylcarbamoyl)benzyl]-4-oxo-1-phenyl-1,4-dihydro-[1,8]naphthyridine-2-carboxylic acid methyl ester (4) was identified as a novel, druglike and selective quinolone pan JNK inhibitor. In this communication, some of the structure-activity relationship of the azaquinolone analogues leading to 4 is discussed. The focus is on how changes at the amide functionality affected the biochemical potency, cellular potency, metabolic properties, and solubility of this class of JNK inhibitors. Optimization of these properties led to the identification of the adamantyl analogue, 4. 4 achieved proof of mechanism in both rat and mouse TNF-α challenge models.
- Hoffmann-La Roche Inc. , pRED, Pharma Research & Early Development, DTA Metabolism, 340 Kingsland Street, Nutley, New Jersey 07110, United States.
Organizational Affiliation: