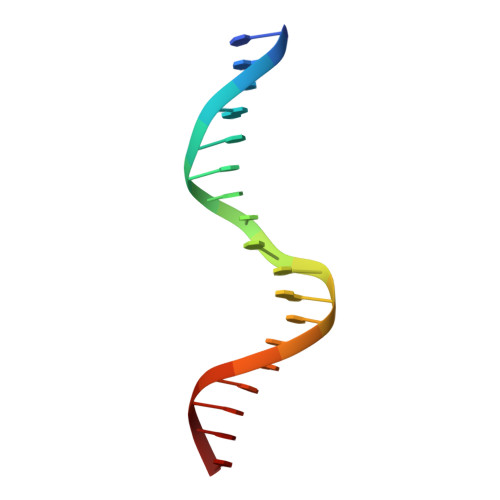
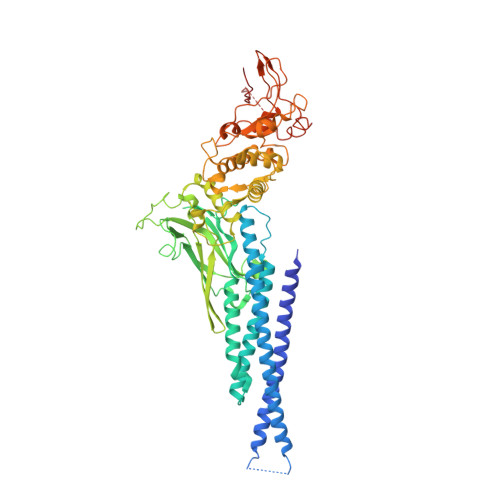Observation of unphosphorylated STAT3 core protein binding to target dsDNA by PEMSA and X-ray crystallography.
Nkansah, E., Shah, R., Collie, G.W., Parkinson, G.N., Palmer, J., Rahman, K.M., Bui, T.T., Drake, A.F., Husby, J., Neidle, S., Zinzalla, G., Thurston, D.E., Wilderspin, A.F.(2013) FEBS Lett 587: 833-839
- PubMed: 23434585 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2013.01.065
- Primary Citation Related Structures:
4E68 - PubMed Abstract:
The STAT3 transcription factor plays a central role in a wide range of cancer types where it is over-expressed. Previously, phosphorylation of this protein was thought to be a prerequisite for direct binding to DNA. However, we have now shown complete binding of a purified unphosphorylated STAT3 (uSTAT3) core directly to M67 DNA, the high affinity STAT3 target DNA sequence, by a protein electrophoretic mobility shift assay (PEMSA). Binding to M67 DNA was inhibited by addition of increasing concentrations of a phosphotyrosyl peptide. X-ray crystallography demonstrates one mode of binding that is similar to that known for the STAT3 core phosphorylated at Y705.
- Department of Pharmaceutical and Biological Chemistry, UCL School of Pharmacy, London, United Kingdom.
Organizational Affiliation: