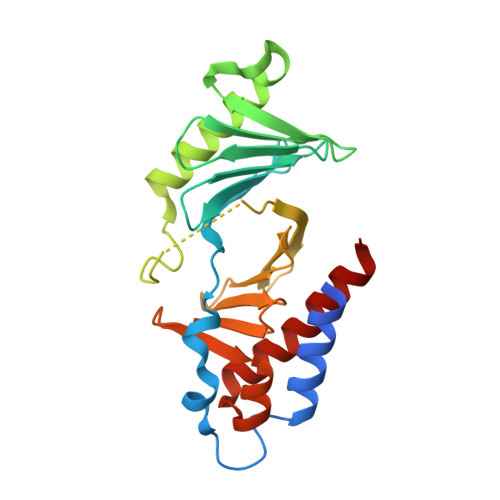
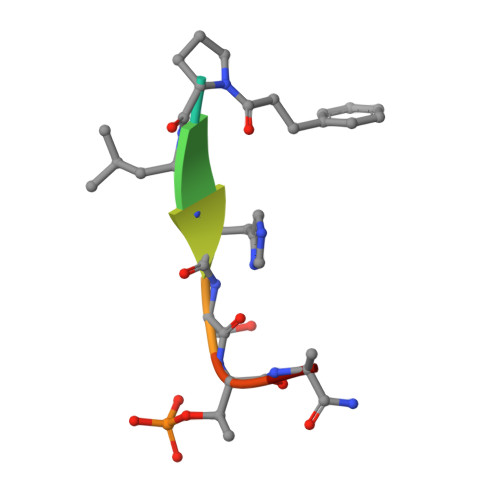
Using ligand-mapping simulations to design a ligand selectively targeting a cryptic surface pocket of polo-like kinase 1.
Tan, Y.S., Sledz, P., Lang, S., Stubbs, C.J., Spring, D.R., Abell, C., Best, R.B.(2012) Angew Chem Int Ed Engl 51: 10078-10081
- PubMed: 22961729 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.201205676
- Primary Citation Related Structures:
4E67 - Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge, CB2 1EW UK.
Organizational Affiliation: