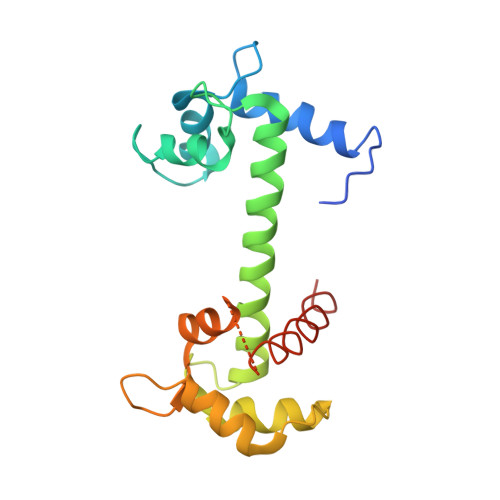
Structural basis for the interaction of unstructured neuron specific substrates neuromodulin and neurogranin with calmodulin
Kumar, V., Chichili, V.P.R., Zhong, L., Tang, X., Velazquez-Campoy, A., Sheu, F.-S., Seetharaman, J., Gerges, N.Z., Sivaraman, J.(2013) Sci Rep 3: 1392-1392
- PubMed: 23462742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep01392
- Primary Citation Related Structures:
4E50, 4E53 - PubMed Abstract:
Neuromodulin (Nm) and neurogranin (Ng) are neuron-specific substrates of protein kinase C (PKC). Their interactions with Calmodulin (CaM) are crucial for learning and memory formation in neurons. Here, we report the structure of IQ peptides (24aa) of Nm/Ng complexed with CaM and their functional studies with full-length proteins. Nm/Ng and their respective IQ peptides are intrinsically unstructured; however, upon binding with CaM, IQ motifs adopt a helical conformation. Ser41 (Ser36) of Nm (Ng) is located in a negatively charged pocket in the apo CaM and, when phosphorylated, it will repel Nm/Ng from CaM. These observations explain the mechanism by which PKC-induced Ser phosphorylation blocks the association of Nm/Ng with CaM and interrupts several learning- and memory-associated functions. Moreover, the present study identified Arg as a key CaM interacting residue from Nm/Ng. This residue is crucial for CaM-mediated function, as evidenced by the inability of the Ng mutant (Arg-to-Ala) to potentiate synaptic transmission in CA1 hippocampal neurons.
- Department of Biological Sciences, National University of Singapore, Singapore.
Organizational Affiliation: