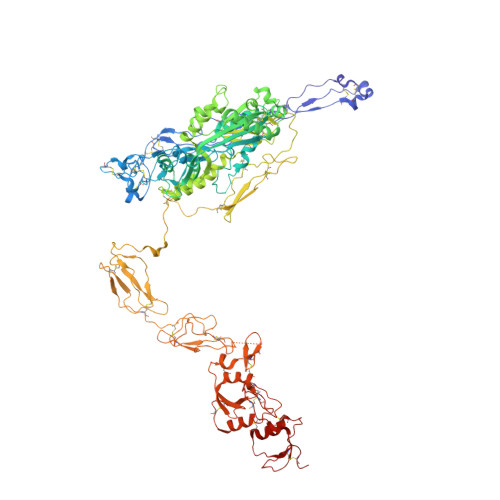
Crystal structure of c5b-6 suggests structural basis for priming assembly of the membrane attack complex.
Aleshin, A.E., Discipio, R.G., Stec, B., Liddington, R.C.(2012) J Biological Chem 287: 19642-19652
- PubMed: 22500023 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.361121
- Primary Citation Related Structures:
4E0S - PubMed Abstract:
The complement membrane attack complex (MAC) forms transmembrane pores in pathogen membranes. The first step in MAC assembly is cleavage of C5 to generate metastable C5b, which forms a stable complex with C6, termed C5b-6. C5b-6 initiates pore formation via the sequential recruitment of homologous proteins: C7, C8, and 12-18 copies of C9, each of which comprises a central MAC-perforin domain flanked by auxiliary domains. We recently proposed a model of pore assembly, in which the auxiliary domains play key roles, both in stabilizing the closed conformation of the protomers and in driving the sequential opening of the MAC-perforin β-sheet of each new recruit to the growing pore. Here, we describe an atomic model of C5b-6 at 4.2 Å resolution. We show that C5b provides four interfaces for the auxiliary domains of C6. The largest interface is created by the insertion of an interdomain linker from C6 into a hydrophobic groove created by a major reorganization of the α-helical domain of C5b. In combination with the rigid body docking of N-terminal elements of both proteins, C5b becomes locked into a stable conformation. Both C6 auxiliary domains flanking the linker pack tightly against C5b. The net effect is to induce the clockwise rigid body rotation of four auxiliary domains, as well as the opening/twisting of the central β-sheet of C6, in the directions predicted by our model to activate or prime C6 for the subsequent steps in MAC assembly. The complex also suggests novel small molecule strategies for modulating pathological MAC assembly.
- Program on Infectious Diseases, Sanford-Burnham Medical Research Institute, La Jolla, California 92037, USA.
Organizational Affiliation: