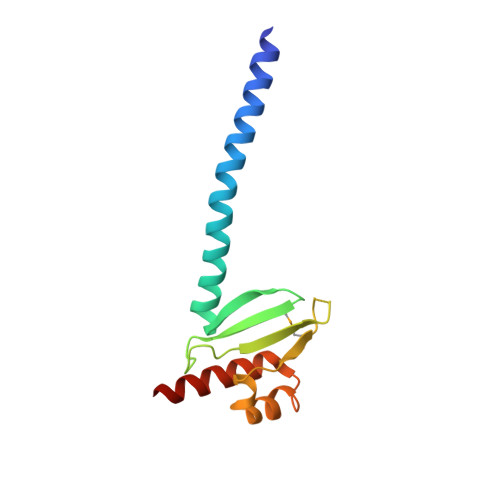
Structure of human Mad1 C-terminal domain reveals its involvement in kinetochore targeting.
Kim, S., Sun, H., Tomchick, D.R., Yu, H., Luo, X.(2012) Proc Natl Acad Sci U S A 109: 6549-6554
- PubMed: 22493223 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1118210109
- Primary Citation Related Structures:
4DZO - PubMed Abstract:
The spindle checkpoint prevents aneuploidy by delaying anaphase onset until all sister chromatids achieve proper microtubule attachment. The kinetochore-bound checkpoint protein complex Mad1-Mad2 promotes the conformational activation of Mad2 and serves as a catalytic engine of checkpoint signaling. How Mad1 is targeted to kinetochores is not understood. Here, we report the crystal structure of the conserved C-terminal domain (CTD) of human Mad1. Mad1 CTD forms a homodimer and, unexpectedly, has a fold similar to those of the kinetochore-binding domains of Spc25 and Csm1. Nonoverlapping Mad1 fragments retain detectable kinetochore targeting. Deletion of the CTD diminishes, does not abolish, Mad1 kinetochore localization. Mutagenesis studies further map the functional interface of Mad1 CTD in kinetochore targeting and implicate Bub1 as its receptor. Our results indicate that CTD is a part of an extensive kinetochore-binding interface of Mad1, and rationalize graded kinetochore targeting of Mad1 during checkpoint signaling.
- Department of Pharmacology, University of Texas Southwestern Medical Center, 6001 Forest Park Road, Dallas, TX 75390, USA.
Organizational Affiliation: