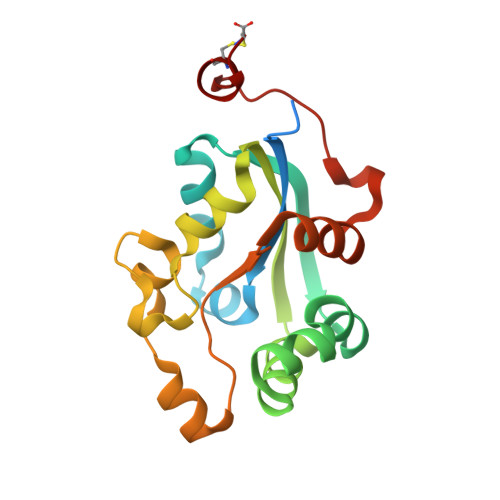
Structure and analysis of nucleoside diphosphate kinase from Borrelia burgdorferi prepared in a transition-state complex with ADP and vanadate moieties.
Dumais, M., Davies, D.R., Lin, T., Staker, B.L., Myler, P.J., Van Voorhis, W.C.(2018) Acta Crystallogr F Struct Biol Commun 74: 373-384
- PubMed: 29870023 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18007392
- Primary Citation Related Structures:
4DI6, 4DZ6 - PubMed Abstract:
Nucleoside diphosphate kinases (NDKs) are implicated in a wide variety of cellular functions owing to their enzymatic conversion of NDP to NTP. NDK from Borrelia burgdorferi (BbNDK) was selected for functional and structural analysis to determine whether its activity is required for infection and to assess its potential for therapeutic inhibition. The Seattle Structural Genomics Center for Infectious Diseases (SSGCID) expressed recombinant BbNDK protein. The protein was crystallized and structures were solved of both the apoenzyme and a liganded form with ADP and vanadate ligands. This provided two structures and allowed the elucidation of changes between the apo and ligand-bound enzymes. Infectivity studies with ndk transposon mutants demonstrated that NDK function was important for establishing a robust infection in mice, and provided a rationale for therapeutic targeting of BbNDK. The protein structure was compared with other NDK structures found in the Protein Data Bank and was found to have similar primary, secondary, tertiary and quaternary structures, with conserved residues acting as the catalytic pocket, primarily using His132 as the phosphohistidine-transfer residue. Vanadate and ADP complexes model the transition state of this phosphoryl-transfer reaction, demonstrating that the pocket closes when bound to ADP, while allowing the addition or removal of a γ-phosphate. This analysis provides a framework for the design of potential therapeutics targeting BbNDK inhibition.
- Department of Allergy and Infectious Disease, University of Washington, Seattle, Washington, USA.
Organizational Affiliation: