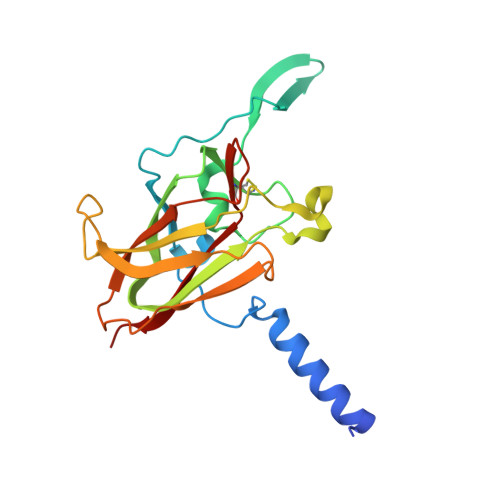
LINC Complexes Form by Binding of Three KASH Peptides to Domain Interfaces of Trimeric SUN Proteins.
Sosa, B.A., Rothballer, A., Kutay, U., Schwartz, T.U.(2012) Cell 149: 1035-1047
- PubMed: 22632968
- DOI: https://doi.org/10.1016/j.cell.2012.03.046
- Primary Citation of Related Structures:
4DXR, 4DXS, 4DXT - PubMed Abstract:
Linker of nucleoskeleton and cytoskeleton (LINC) complexes span the nuclear envelope and are composed of KASH and SUN proteins residing in the outer and inner nuclear membrane, respectively. LINC formation relies on direct binding of KASH and SUN in the perinuclear space. Thereby, molecular tethers are formed that can transmit forces for chromosome movements, nuclear migration, and anchorage. We present crystal structures of the human SUN2-KASH1/2 complex, the core of the LINC complex. The SUN2 domain is rigidly attached to a trimeric coiled coil that prepositions it to bind three KASH peptides. The peptides bind in three deep and expansive grooves formed between adjacent SUN domains, effectively acting as molecular glue. In addition, a disulfide between conserved cysteines on SUN and KASH covalently links both proteins. The structure provides the basis of LINC complex formation and suggests a model for how LINC complexes might arrange into higher-order clusters to enhance force-coupling.
- Department of Biology, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, MA 02139, USA.
Organizational Affiliation: