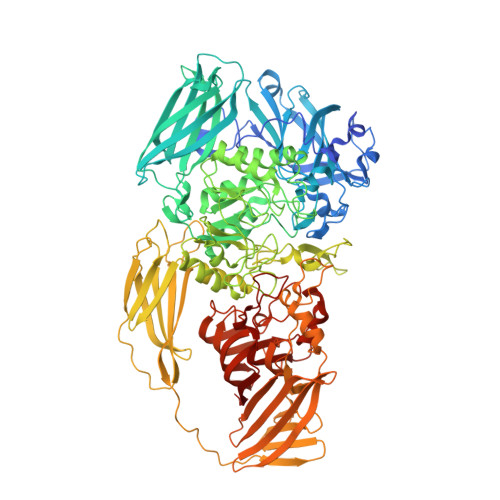
Structural Explanation for Allolactose (lac operon inducer) Synthesis by lacZ beta-Galactosidase and the Evolutionary Relationship between Allolactose synthesis and the lac Repressor
Wheatley, R.W., Lo, S., Jancewicz, L.J., Dugdale, M.L., Huber, R.E.(2013) J Biological Chem 288: 12993-13005
- PubMed: 23486479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.455436
- Primary Citation Related Structures:
4DUW, 4DUX - PubMed Abstract:
β-Galactosidase (lacZ) has bifunctional activity. It hydrolyzes lactose to galactose and glucose and catalyzes the intramolecular isomerization of lactose to allolactose, the lac operon inducer. β-Galactosidase promotes the isomerization by means of an acceptor site that binds glucose after its cleavage from lactose and thus delays its exit from the site. However, because of its relatively low affinity for glucose, details of this site have remained elusive. We present structural data mapping the glucose site based on a substituted enzyme (G794A-β-galactosidase) that traps allolactose. Various lines of evidence indicate that the glucose of the trapped allolactose is in the acceptor position. The evidence includes structures with Bis-Tris (2,2-bis(hydroxymethyl)-2,2',2″-nitrilotriethanol) and L-ribose in the site and kinetic binding studies with substituted β-galactosidases. The site is composed of Asn-102, His-418, Lys-517, Ser-796, Glu-797, and Trp-999. Ser-796 and Glu-797 are part of a loop (residues 795-803) that closes over the active site. This loop appears essential for the bifunctional nature of the enzyme because it helps form the glucose binding site. In addition, because the loop is mobile, glucose binding is transient, allowing the release of some glucose. Bioinformatics studies showed that the residues important for interacting with glucose are only conserved in a subset of related enzymes. Thus, intramolecular isomerization is not a universal feature of β-galactosidases. Genomic analyses indicated that lac repressors were co-selected only within the conserved subset. This shows that the glucose binding site of β-galactosidase played an important role in lac operon evolution.
- Division of Biochemistry, Faculty of Science, University of Calgary, Calgary, Alberta T2N 1N4, Canada.
Organizational Affiliation: