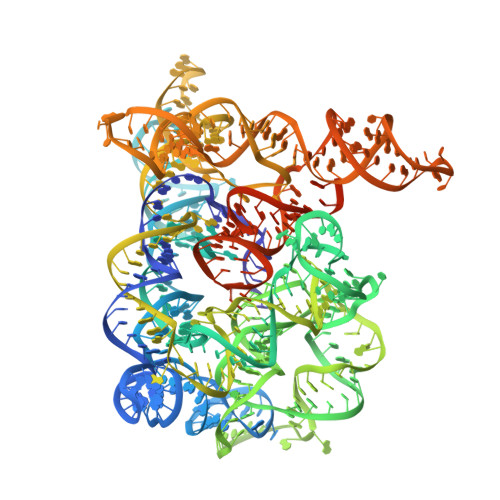
Crystal structure of a group II intron in the pre-catalytic state.
Chan, R.T., Robart, A.R., Rajashankar, K.R., Pyle, A.M., Toor, N.(2012) Nat Struct Mol Biol 19: 555-557
- PubMed: 22484319 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2270
- Primary Citation Related Structures:
4DS6 - PubMed Abstract:
Group II introns are self-splicing catalytic RNAs that are thought to be ancestral to the spliceosome. Here we report the 3.65-Å crystal structure of the group II intron from Oceanobacillus iheyensis in the pre-catalytic state. The structure reveals the conformation of the 5' splice site in the catalytic core and represents the first structure of an intron prior to the first step of splicing.
- Department of Chemistry and Biochemistry, University of California, San Diego, San Diego, California, USA.
Organizational Affiliation: