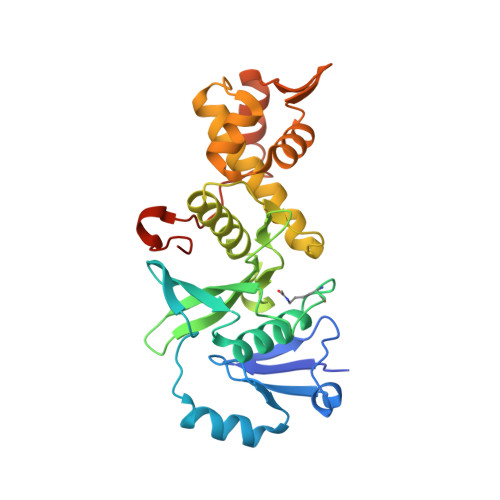
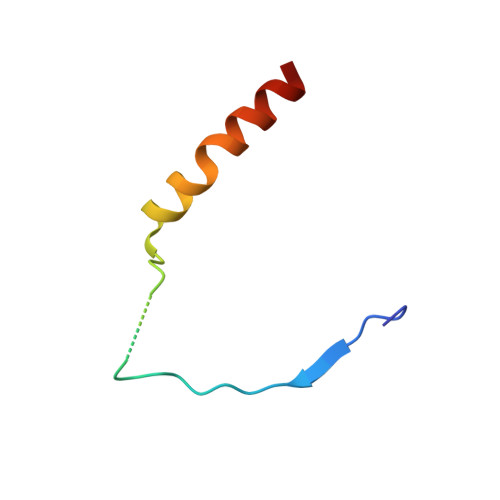
Structural insight into the regulation of MOF in the male-specific lethal complex and the non-specific lethal complex.
Huang, J., Wan, B., Wu, L., Yang, Y., Dou, Y., Lei, M.(2012) Cell Res 22: 1078-1081
- PubMed: 22547026 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2012.72
- Primary Citation Related Structures:
4DNC