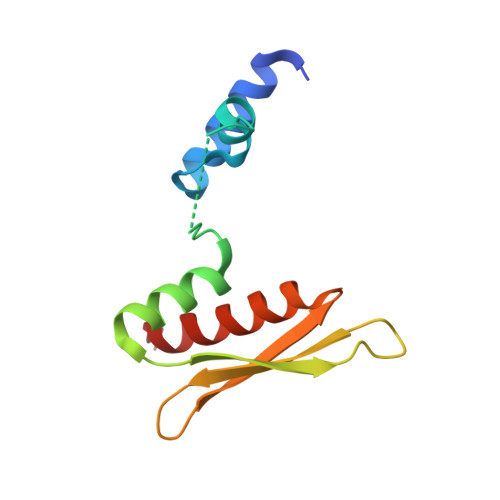
Staufen1 dimerizes through a conserved motif and a degenerate dsRNA-binding domain to promote mRNA decay.
Gleghorn, M.L., Gong, C., Kielkopf, C.L., Maquat, L.E.(2013) Nat Struct Mol Biol 20: 515-524
- PubMed: 23524536 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2528
- Primary Citation Related Structures:
4DKK - PubMed Abstract:
Staufen1 (STAU1)-mediated mRNA decay (SMD) degrades mammalian-cell mRNAs that bind the double-stranded RNA (dsRNA)-binding protein STAU1 in their 3' untranslated region. We report a new motif, which typifies STAU homologs from all vertebrate classes, that is responsible for human STAU1 (hSTAU1) homodimerization. Our crystal structure and mutagenesis analyses reveal that this motif, which we named the Staufen-swapping motif (SSM), and the dsRNA-binding domain 5 ('RBD'5) mediate protein dimerization: the two SSM α-helices of one molecule interact primarily through a hydrophobic patch with the two 'RBD'5 α-helices of a second molecule. 'RBD'5 adopts the canonical α-β-β-β-α fold of a functional RBD, but it lacks residues and features required to bind duplex RNA. In cells, SSM-mediated hSTAU1 dimerization increases the efficiency of SMD by augmenting hSTAU1 binding to the ATP-dependent RNA helicase hUPF1. Dimerization regulates keratinocyte-mediated wound healing and many other cellular processes.
- Department of Biochemistry and Biophysics, School of Medicine and Dentistry, University of Rochester, Rochester, New York, USA.
Organizational Affiliation: