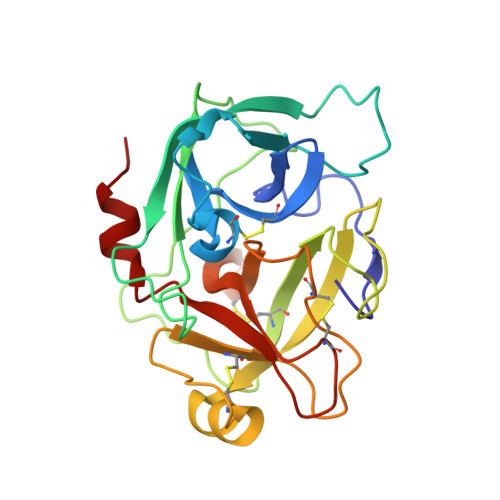
Crystal structure of a supercharged variant of the human enteropeptidase light chain.
Simeonov, P., Zahn, M., Strater, N., Zuchner, T.(2012) Proteins 80: 1907-1910
- PubMed: 22488687 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24084
- Primary Citation Related Structures:
4DGJ - PubMed Abstract:
The highly specific serine protease human enteropeptidase light chain cleaves the Asp4Lys recognition sequence and represents an interesting enzyme for biotechnological applications. The human enzyme shows 10 times faster kinetics compared to other animal sources but low solubility under low salt conditions, which hampers protein production and crystallization. Therefore, a supercharged variant (N6D/G21D/G22D/N142D/K210E/C112S) with increased solubility was used for crystallization. The structure (resolution, 1.9 Å) displays a typical α/β trypsin-like serine protease-fold. The mutations introduced for protein supercharging generate larger clusters of negative potential on both sites of the active cleft but do not affect the structural integrity of the protein.
- Institute of Bioanalytical Chemistry, Center for Biotechnology and Biomedicine, Universität Leipzig, 04103 Leipzig, Germany.
Organizational Affiliation: