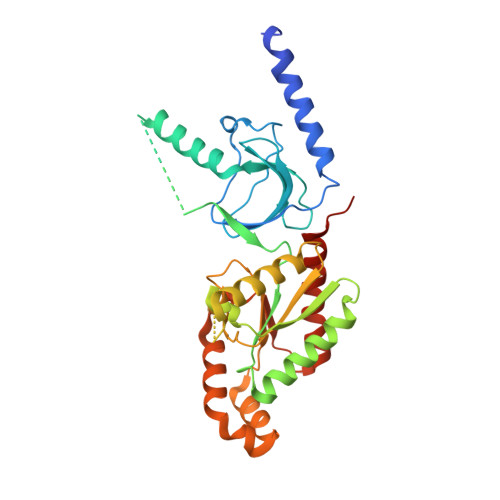
The role of a voltage-dependent Ca2+ channel intracellular linker: a structure-function analysis.
Almagor, L., Chomsky-Hecht, O., Ben-Mocha, A., Hendin-Barak, D., Dascal, N., Hirsch, J.A.(2012) J Neurosci 32: 7602-7613
- PubMed: 22649239 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1523/JNEUROSCI.5727-11.2012
- Primary Citation Related Structures:
4DEX, 4DEY - PubMed Abstract:
Voltage-dependent calcium channels (VDCCs) allow the passage of Ca(2+) ions through cellular membranes in response to membrane depolarization. The channel pore-forming subunit, α1, and a regulatory subunit (Ca(V)β) form a high affinity complex where Ca(V)β binds to a α1 interacting domain in the intracellular linker between α1 membrane domains I and II (I-II linker). We determined crystal structures of Ca(V)β2 functional core in complex with the Ca(V)1.2 and Ca(V)2.2 I-II linkers to a resolution of 1.95 and 2.0 Å, respectively. Structural differences between the highly conserved linkers, important for coupling Ca(V)β to the channel pore, guided mechanistic functional studies. Electrophysiological measurements point to the importance of differing linker structure in both Ca(V)1 and 2 subtypes with mutations affecting both voltage- and calcium-dependent inactivation and voltage dependence of activation. These linker effects persist in the absence of Ca(V)β, pointing to the intrinsic role of the linker in VDCC function and suggesting that I-II linker structure can serve as a brake during inactivation.
- Departments of Biochemistry and Molecular Biology, Institute of Structural Biology, George S. Wise Faculty of Life Sciences, Sackler Faculty of Medicine, Tel Aviv University, Tel Aviv 69978, Israel.
Organizational Affiliation: