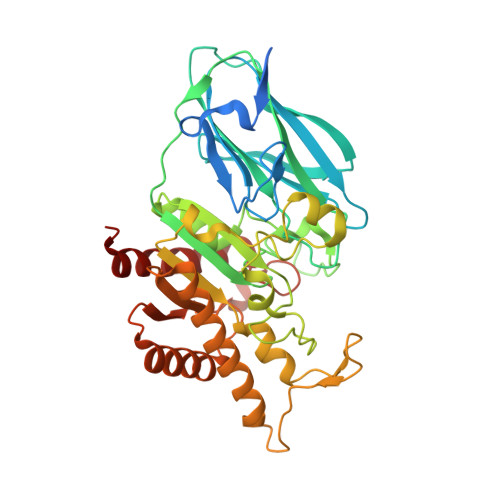
Structure and function of an acetyl xylan esterase (Est2A) from the rumen bacterium Butyrivibrio proteoclasticus.
Till, M., Goldstone, D.C., Attwood, G.T., Moon, C.D., Kelly, W.J., Arcus, V.L.(2013) Proteins 81: 911-917
- PubMed: 23345031 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24254
- Primary Citation Related Structures:
3U37, 4DEV - PubMed Abstract:
Butyrivibrio proteoclasticus is a significant component of the microbial population of the rumen of dairy cattle. It is a xylan-degrading organism whose genome encodes a large number of open reading frames annotated as fiber-degrading enzymes. We have determined the three-dimensional structure of Est2A, an acetyl xylan esterase from B. proteoclasticus, at 2.1 Å resolution, along with the structure of an inactive mutant (H351A) at 2.0 Å resolution. The structure reveals two domains-a C-terminal SGNH domain and an N-terminal jelly-roll domain typical of CE2 family structures. The structures are accompanied by experimentally determined enzymatic parameters against two model substrates, para-nitrophenyl acetate and para-nitrophenyl butyrate. The suite of fiber-degrading enzymes produced by B. proteoclasticus provides a rich source of new enzymes of potential use in industrial settings.
- Department of Biological Sciences, University of Waikato, Hamilton, New Zealand.
Organizational Affiliation: