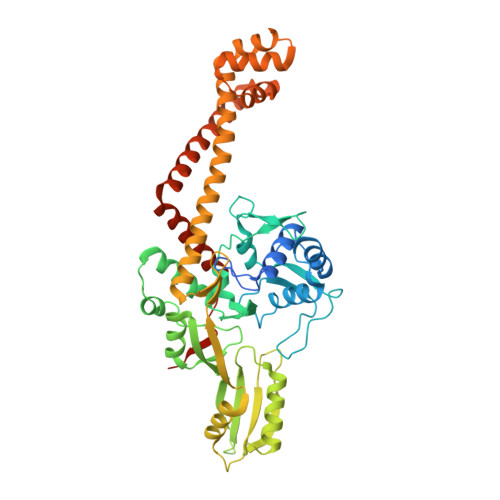
Mapping the Spectrum of Conformational States of the DNA- and C-Gates in Bacillus subtilis Gyrase.
Rudolph, M.G., Klostermeier, D.(2013) J Mol Biology 425: 2632-2640
- PubMed: 23602808 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.04.010
- Primary Citation Related Structures:
4DDQ - PubMed Abstract:
Type II DNA topoisomerases alter the supercoiling state of DNA in an ATP-dependent fashion that requires large conformational changes. The directionality of DNA strand transfer is controlled by three transient protein interfaces, termed the N-gate, DNA-gate, and C-gate. Bacterial gyrase is a type II DNA topoisomerase of A2B2 composition. The N-gate is formed by the two GyrB subunits and the GyrA subunits form the DNA- and C-gates. In structures of type II topoisomerase fragments, the DNA- and C-gates delimit a cavity for DNA and can be open or closed. However, the conformational space accessible has not yet been mapped. Here, we describe the crystal structure of the Bacillus subtilis DNA gyrase A subunit lacking the C-terminal DNA-wrapping domains. Five dimeric states of the GyrA N-terminal domain are observed, with their DNA- and C-gates either closed, or open to different extents. All of these conformations can in principle accommodate double-stranded DNA in the central cavity but only one conformation has its DNA-gate open wide enough for DNA to enter. The structure thus reflects the lower limit of DNA-gate opening that must occur during gyrase catalysis. The DNA-gate is formed by two flat surfaces, with few interactions. In contrast, the C-gate exhibits a highly undulated surface and forms a large number of interactions. None of the dimers in the crystal structures display an open C-gate that would allow DNA passage, in agreement with a transient opening of this gate during the catalytic cycle of DNA supercoiling.
- F. Hoffmann-La Roche AG, Pharma Research and Early Development, Discovery Technologies, Grenzacherstrasse 124, CH-4070 Basel, Switzerland. dagmar.klostermeier@uni-muenster.de
Organizational Affiliation: