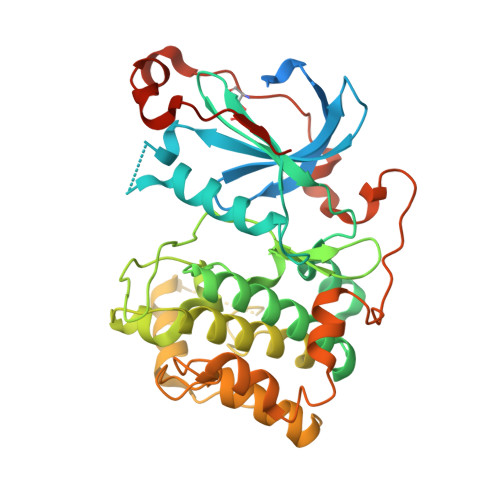
Substrate recognition mechanism of atypical protein kinase Cs revealed by the structure of PKC iota in complex with a substrate peptide from Par-3
Wang, C., Shang, Y., Yu, J., Zhang, M.(2012) Structure 20: 791-801
- PubMed: 22579248 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.02.022
- Primary Citation Related Structures:
4DC2 - PubMed Abstract:
Protein kinase C (PKC) play critical roles in many cellular functions including differentiation, proliferation, growth, and survival. However, the molecular bases governing PKC's substrate recognitions remain poorly understood. Here we determined the structure of PKCι in complex with a peptide from Par-3 at 2.4 Å. PKCι in the complex adopts catalytically competent, closed conformation without phosphorylation of Thr402 in the activation loop. The Par-3 peptide binds to an elongated groove formed by the N- and C-lobes of the kinase domain. The PKCι/Par-3 complex structure, together with extensive biochemical studies, reveals a set of substrate recognition sites common to all PKC isozymes as well as a hydrophobic pocket unique to aPKC. A consensus aPKC's substrate recognition sequence pattern can be readily identified based on the complex structure. Finally, we demonstrate that the pseudosubstrate sequence of PKCι resembles its substrate sequence, directly binds to and inhibits the activity of the kinase.
- Division of Life Science, State Key Laboratory of Molecular Neuroscience, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong.
Organizational Affiliation: