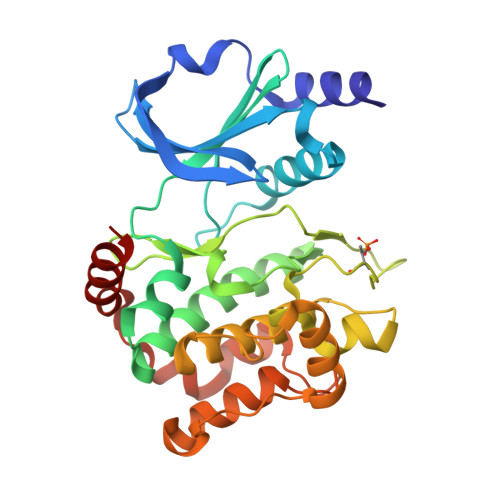
The art of filling protein pockets efficiently with octahedral metal complexes.
Blanck, S., Maksimoska, J., Baumeister, J., Harms, K., Marmorstein, R., Meggers, E.(2012) Angew Chem Int Ed Engl 51: 5244-5246
- PubMed: 22383326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.201108865
- Primary Citation Related Structures:
4DAW - Fachbereich Chemie, Philipps-Universität Marburg, Hans-Meerwein-Strasse, 35043 Marburg, Germany.
Organizational Affiliation: