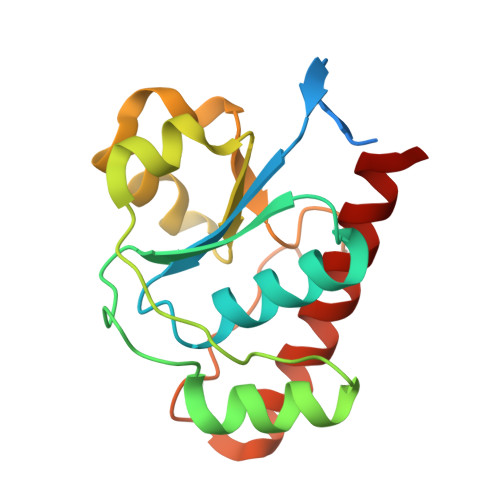
Characterization and 1.57 A Resolution Structure of the Key Fire Blight Phosphatase Amsi from Erwinia Amylovora
Salomone-Stagni, M., Musiani, F., Benini, S.(2016) Acta Crystallogr Sect F Struct Biol Cryst Commun 72: 903
- PubMed: 27917839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16018781
- Primary Citation Related Structures:
4D74 - PubMed Abstract:
AmsI is a low-molecular-weight protein tyrosine phosphatase that regulates the production of amylovoran in the Gram-negative bacterium Erwinia amylovora, a specific pathogen of rosaceous plants such as apple, pear and quince. Amylovoran is an exopolysaccharide that is necessary for successful infection. In order to shed light on AmsI, its structure was solved at 1.57 Å resolution at the same pH as its highest measured activity (pH 5.5). In the active site, a water molecule, bridging between the catalytic Arg15 and the reaction-product analogue sulfate, might be representative of the water molecule attacking the phospho-cysteine intermediate in the second step of the reaction mechanism.
- Faculty of Science and Technology, Free University of Bozen-Bolzano, Piazza Università 5, 39100 Bolzano, Italy.
Organizational Affiliation: