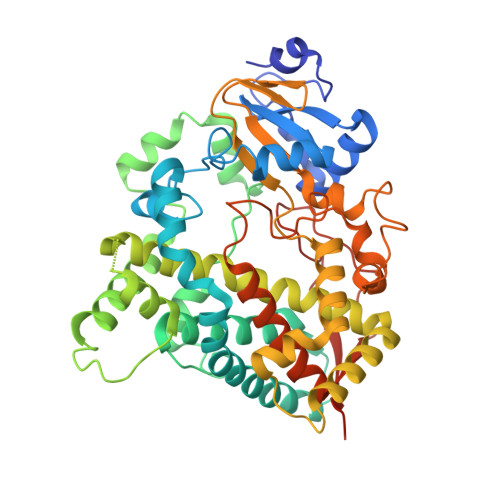
Structure-Based Inhibitor Design for Evaluation of a Cyp3A4 Pharmacophore Model.
Kaur, P., Chamberlin, R., Poulos, T.L., Sevrioukova, I.F.(2016) J Med Chem 59: 4210
- PubMed: 26371436 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01146
- Primary Citation Related Structures:
4D6Z, 4D75, 4D78, 4D7D - PubMed Abstract:
Human cytochrome P450 3A4 (CYP3A4) is a key xenobiotic-metabolizing enzyme that oxidizes and clears the majority of drugs. CYP3A4 inhibition may lead to drug-drug interactions, toxicity, and other adverse effects but, in some cases, could be beneficial and enhance therapeutic efficiency of coadministered pharmaceuticals that are metabolized by CYP3A4. On the basis of our investigations of analogs of ritonavir, a potent CYP3A4 inactivator and pharmacoenhancer, we have built a pharmacophore model for a CYP3A4-specific inhibitor. This study is the first attempt to test this model using a set of rationally designed compounds. The functional and structural data presented here agree well with the proposed pharmacophore. In particular, we confirmed the importance of a flexible backbone, the H-bond donor/acceptor moiety, and aromaticity of the side group analogous to Phe-2 of ritonavir and demonstrated the leading role of hydrophobic interactions at the sites adjacent to the heme and phenylalanine cluster in the ligand binding process. The X-ray structures of CYP3A4 bound to the rationally designed inhibitors provide deeper insights into the mechanism of the CYP3A4-ligand interaction. Most importantly, two of our compounds (15a and 15b) that are less complex than ritonavir have comparable submicromolar affinity and inhibitory potency for CYP3A4 and, thus, could serve as templates for synthesis of second generation inhibitors for further evaluation and optimization of the pharmacophore model.
- Departments of †Pharmaceutical Sciences, ‡Chemistry, and §Molecular Biology and Biochemistry, University of California-Irvine , Irvine, California 92697, United States.
Organizational Affiliation: