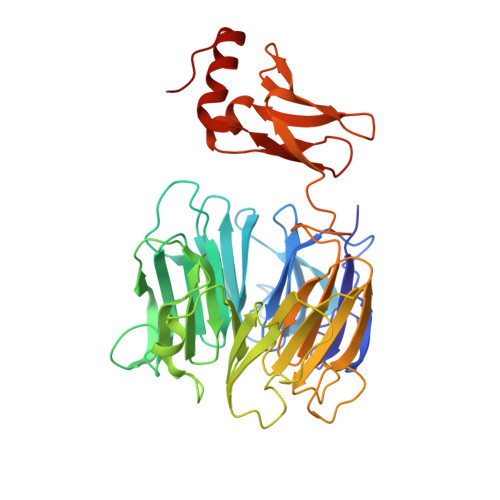
Structure of the Kti11/Kti13 Heterodimer and its Double Role in Modifications of tRNA and Eukaryotic Elongation Factor 2.
Glatt, S., Zabel, R., Vonkova, I., Kumar, A., Netz, D.J., Pierik, A.J., Rybin, V., Lill, R., Gavin, A., Balbach, J., Breunig, K.D., Muller, C.W.(2015) Structure 23: 7
- PubMed: 25543256
- DOI: https://doi.org/10.1016/j.str.2014.11.008
- Primary Citation of Related Structures:
4D4O, 4D4P, 4D4Q - PubMed Abstract:
The small, highly conserved Kti11 alias Dph3 protein encoded by the Kluyveromyces lactis killer toxin insensitive gene KTI11/DPH3 is involved in the diphthamide modification of eukaryotic elongation factor 2 and, together with Kti13, in Elongator-dependent tRNA wobble base modifications, thereby affecting the speed and accuracy of protein biosynthesis through two distinct mechanisms. We have solved the crystal structures of Saccharomyces cerevisiae Kti13 and the Kti11/Kti13 heterodimer at 2.4 and 2.9 Å resolution, respectively, and validated interacting residues through mutational analysis in vitro and in vivo. We show that metal coordination by Kti11 and its heterodimerization with Kti13 are essential for both translational control mechanisms. Our structural and functional analyses identify Kti13 as an additional component of the diphthamide modification pathway and provide insight into the molecular mechanisms that allow the Kti11/Kti13 heterodimer to coregulate two consecutive steps in ribosomal protein synthesis.
- Structural and Computational Biology Unit, European Molecular Biology Laboratory, Meyerhofstraße 1, 69117 Heidelberg, Germany.
Organizational Affiliation: