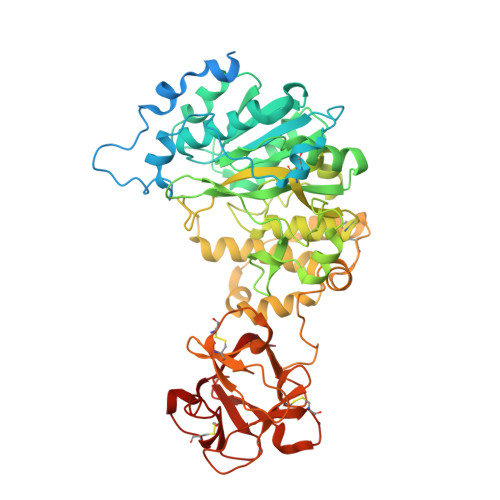
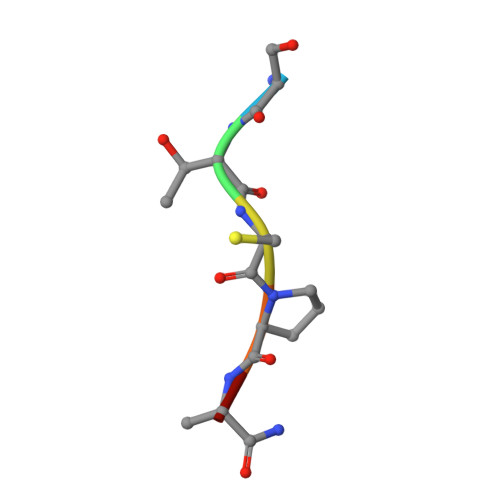
Substrate-Guided Front-Face Reaction Revealed by Combined Structural Snapshots and Metadynamics for the Polypeptide N- Acetylgalactosaminyltransferase 2.
Lira-Navarrete, E., Iglesias-Fernandez, J., Zandberg, W.F., Companon, I., Kong, Y., Corzana, F., Pinto, B.M., Clausen, H., Peregrina, J.M., Vocadlo, D., Rovira, C., Hurtado-Guerrero, R.(2014) Angew Chem Int Ed Engl 53: 8206
- PubMed: 24954443
- DOI: https://doi.org/10.1002/anie.201402781
- Primary Citation of Related Structures:
4D0T, 4D0Z, 4D11 - PubMed Abstract:
The retaining glycosyltransferase GalNAc-T2 is a member of a large family of human polypeptide GalNAc-transferases that is responsible for the post-translational modification of many cell-surface proteins. By the use of combined structural and computational approaches, we provide the first set of structural snapshots of the enzyme during the catalytic cycle and combine these with quantum-mechanics/molecular-mechanics (QM/MM) metadynamics to unravel the catalytic mechanism of this retaining enzyme at the atomic-electronic level of detail. Our study provides a detailed structural rationale for an ordered bi-bi kinetic mechanism and reveals critical aspects of substrate recognition, which dictate the specificity for acceptor Thr versus Ser residues and enforce a front-face SN i-type reaction in which the substrate N-acetyl sugar substituent coordinates efficient glycosyl transfer.
- Institute of Biocomputation and Physics of Complex Systems (BIFI), University of Zaragoza, BIFI-IQFR (CSIC) Joint Unit, Mariano Esquillor s/n, Campus Rio Ebro, Edificio I+D, Fundacion ARAID, Edificio Pignatelli 36 (Spain).
Organizational Affiliation: