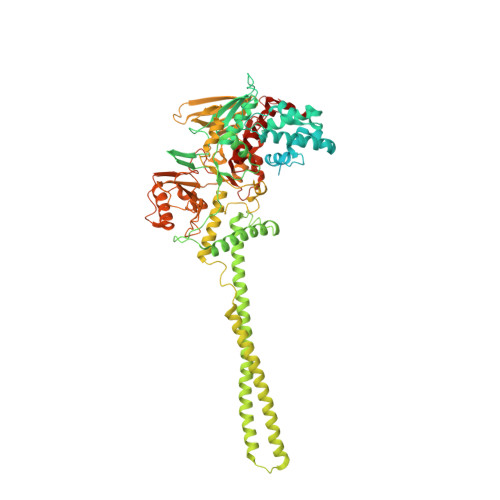
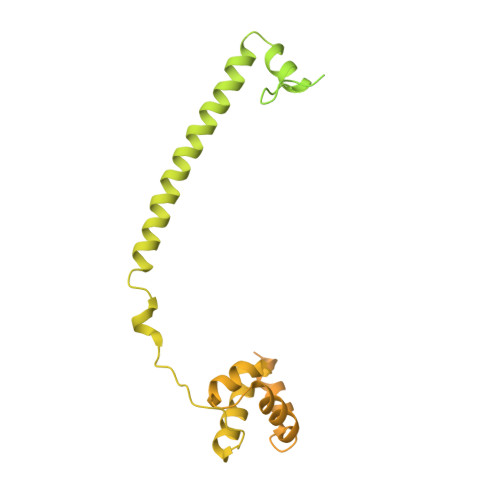
Differential Properties of Transcriptional Complexes Formed by the Corest Family.
Barrios, A.P., Gomez, A.V., Saez, J.E., Ciossani, G., Toffolo, E., Battaglioli, E., Mattevi, A., Andres, M.E.(2014) Mol Cell Biol 34: 2760
- PubMed: 24820421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.00083-14
- Primary Citation Related Structures:
4CZZ - PubMed Abstract:
Mammalian genomes harbor three CoREST genes. rcor1 encodes CoREST (CoREST1), and the paralogues rcor2 and rcor3 encode CoREST2 and CoREST3, respectively. Here, we describe specific properties of transcriptional complexes formed by CoREST proteins with the histone demethylase LSD1/KDM1A and histone deacetylases 1 and 2 (HDAC1/2) and the finding that all three CoRESTs are expressed in the adult rat brain. CoRESTs interact equally strongly with LSD1/KDM1A. Structural analysis shows that the overall conformation of CoREST3 is similar to that of CoREST1 complexed with LSD1/KDM1A. Nonetheless, transcriptional repressive capacity of CoREST3 is lower than that of CoREST1, which correlates with the observation that CoREST3 leads to a reduced LSD1/KDM1A catalytic efficiency. Also, CoREST2 shows a lower transcriptional repression than CoREST1, which is resistant to HDAC inhibitors. CoREST2 displays lower interaction with HDAC1/2, which is barely present in LSD1/KDM1A-CoREST2 complexes. A nonconserved leucine in the first SANT domain of CoREST2 severely weakens its association with HDAC1/2. Furthermore, CoREST2 mutants with increased HDAC1/2 interaction and those without HDAC1/2 interaction exhibit equivalent transcriptional repression capacities, indicating that CoREST2 represses in an HDAC-independent manner. In conclusion, differences among CoREST proteins are instrumental in the modulation of protein-protein interactions and catalytic activities of LSD1/KDM1A-CoREST-HDAC complexes, fine-tuning gene expression regulation.