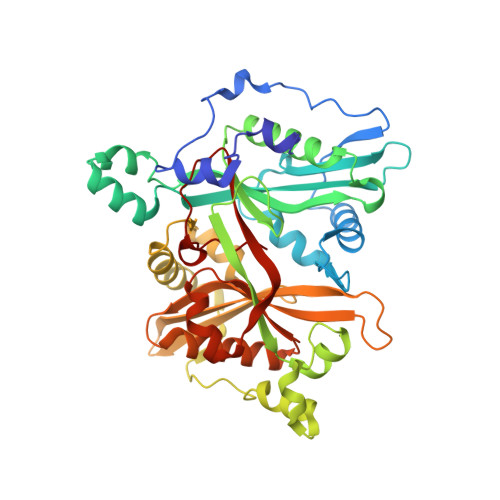
Structure-Based Design of Potent and Selective Leishmania N- Myristoyltransferase Inhibitors.
Hutton, J.A., Goncalves, V., Brannigan, J.A., Paape, D., Wright, M.H., Waugh, T.M., Roberts, S.M., Bell, A.S., Wilkinson, A.J., Smith, D.F., Leatherbarrow, R.J., Tate, E.W.(2014) J Med Chem 57: 8664
- PubMed: 25238611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm5011397
- Primary Citation Related Structures:
4CYN, 4CYO, 4CYP, 4CYQ - PubMed Abstract:
Inhibitors of Leishmania N-myristoyltransferase (NMT), a potential target for the treatment of leishmaniasis, obtained from a high-throughput screen, were resynthesized to validate activity. Crystal structures bound to Leishmania major NMT were obtained, and the active diastereoisomer of one of the inhibitors was identified. On the basis of structural insights, enzyme inhibition was increased 40-fold through hybridization of two distinct binding modes, resulting in novel, highly potent Leishmania donovani NMT inhibitors with good selectivity over the human enzyme.
- Department of Chemistry, Imperial College London , London SW7 2AZ, U.K.
Organizational Affiliation: