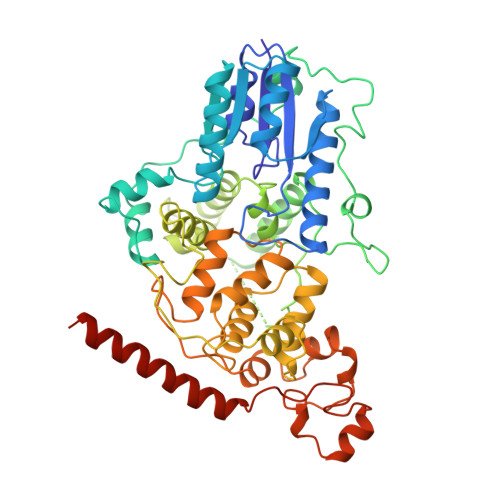
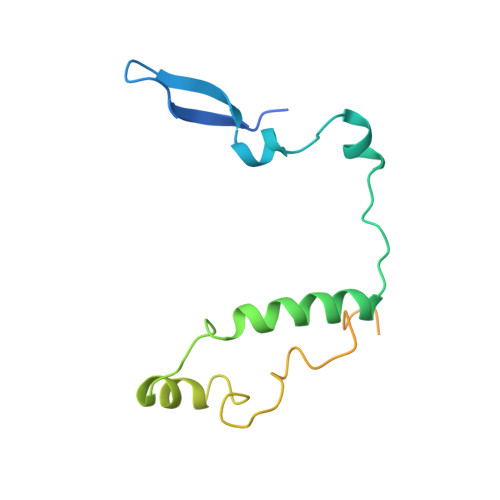
Interaction of Circadian Clock Proteins Cry1 and Per2 is Modulated by Zinc Binding and Disulfide Bond Formation.
Schmalen, I., Reischl, S., Wallach, T., Klemz, R., Grudziecki, A., Prabu, J.R., Benda, C., Kramer, A., Wolf, E.(2014) Cell 157: 1203
- PubMed: 24855952 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2014.03.057
- Primary Citation Related Structures:
4CT0 - PubMed Abstract:
Period (PER) proteins are essential components of the mammalian circadian clock. They form complexes with cryptochromes (CRY), which negatively regulate CLOCK/BMAL1-dependent transactivation of clock and clock-controlled genes. To define the roles of mammalian CRY/PER complexes in the circadian clock, we have determined the crystal structure of a complex comprising the photolyase homology region of mouse CRY1 (mCRY1) and a C-terminal mouse PER2 (mPER2) fragment. mPER2 winds around the helical mCRY1 domain covering the binding sites of FBXL3 and CLOCK/BMAL1, but not the FAD binding pocket. Our structure revealed an unexpected zinc ion in one interface, which stabilizes mCRY1-mPER2 interactions in vivo. We provide evidence that mCRY1/mPER2 complex formation is modulated by an interplay of zinc binding and mCRY1 disulfide bond formation, which may be influenced by the redox state of the cell. Our studies may allow for the development of circadian and metabolic modulators.
- Department of Structural Cell Biology, Max Planck Institute of Biochemistry, Am Klopferspitz 18, 82152 Martinsried/Munich, Germany.
Organizational Affiliation: