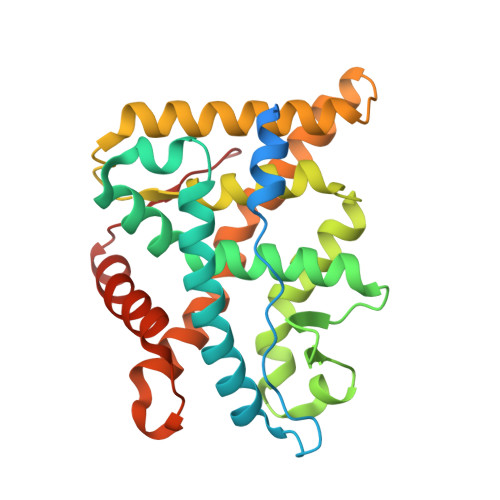
The Discovery of Potent and Selective Non-Steroidal Glucocorticoid Receptor Modulators, Suitable for Inhalation.
Edman, K., Ahlgren, R., Bengtsson, M., Bladh, H., Backstrom, S., Dahmen, J., Henriksson, K., Hillertz, P., Hulikal, V., Jerre, A., Kinchin, L., Kase, C., Lepisto, M., Mile, I., Nilsson, S., Smailagic, A., Taylor, J., Tjornebo, A., Wissler, L., Hansson, T.(2014) Bioorg Med Chem Lett 24: 2571
- PubMed: 24755427 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2014.03.070
- Primary Citation Related Structures:
4CSJ - PubMed Abstract:
We report the discovery of highly potent and selective non-steroidal glucocorticoid receptor modulators with PK properties suitable for inhalation. A high throughput screen of the AstraZeneca compound collection identified sulfonamide 3 as a potent non-steroidal glucocorticoid receptor ligand. Further optimization of this lead generated indazoles 30 and 48 that were progressed to characterization in in vivo models. X-ray crystallography was used to gain further insight into the binding mode of selected ligands.
- Discovery Sciences, Innovative Medicines, AstraZeneca R&D, Pepparedsleden 1, SE-431 83 Mölndal, Sweden.
Organizational Affiliation: