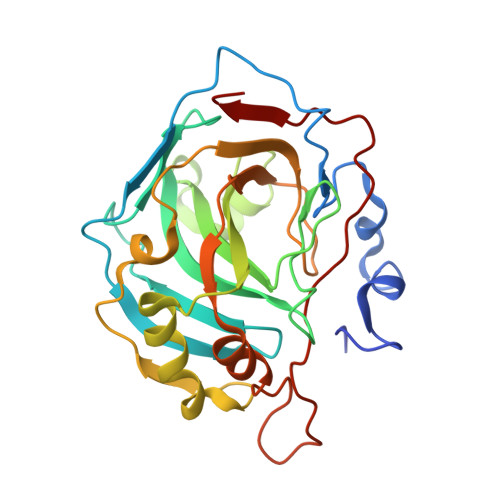
Rational Engineering of a Mesohalophilic Carbonic Anhydrase to an Extreme Halotolerant Biocatalyst.
Warden, A.C., Williams, M., Peat, T.S., Seabrook, S.A., Newman, J., Dojchinov, G., Haritos, V.S.(2015) Nat Commun 6: 10278
- PubMed: 26687908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms10278
- Primary Citation Related Structures:
4CNR, 4CNV, 4CNW, 4CNX, 5A25 - PubMed Abstract:
Enzymes expressed by highly salt-tolerant organisms show many modifications compared with salt-affected counterparts including biased amino acid and lower α-helix content, lower solvent accessibility and negative surface charge. Here, we show that halotolerance can be generated in an enzyme solely by modifying surface residues. Rational design of carbonic anhydrase II is undertaken in three stages replacing 18 residues in total, crystal structures confirm changes are confined to surface residues. Catalytic activities and thermal unfolding temperatures of the designed enzymes increase at high salt concentrations demonstrating their shift to halotolerance, whereas the opposite response is found in the wild-type enzyme. Molecular dynamics calculations reveal a key role for sodium ions in increasing halotolerant enzyme stability largely through interactions with the highly ordered first Na(+) hydration shell. For the first time, an approach to generate extreme halotolerance, a trait with broad application in industrial biocatalysis, in a wild-type enzyme is demonstrated.
- Energy Flagship, Commonwealth Scientific and Industrial Research Organisation (CSIRO), GPO Box 1700, Canberra, Australian Capital Territory 2601, Australia.
Organizational Affiliation: