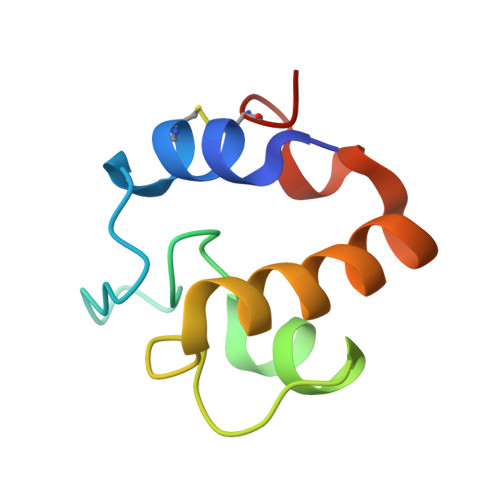
Mycobacterium Tuberculosis Rpfe Crystal Structure Reveals a Positively Charged Catalytic Cleft.
Mavrici, D., Prigozhin, D.M., Alber, T.(2014) Protein Sci 23: 481
- PubMed: 24452911 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2431
- Primary Citation Related Structures:
4CGE - PubMed Abstract:
Resuscitation promoting factor (Rpf) proteins, which hydrolyze the sugar chains in cell-wall peptidoglycan (PG), play key roles in prokaryotic cell elongation, division, and escape from dormancy to vegetative growth. Like other bacteria, Mycobacterium tuberculosis (Mtb) expresses multiple Rpfs, none of which is individually essential. This redundancy has left unclear the distinct functions of the different Rpfs. To explore the distinguishing characteristics of the five Mtb Rpfs, we determined the crystal structure of the RpfE catalytic domain. The protein adopts the characteristic Rpf fold, but the catalytic cleft is narrower compared to Mtb RpfB. Also in contrast to RpfB, in which the substrate-binding surfaces are negatively charged, the corresponding RpfE catalytic pocket and predicted peptide-binding sites are more positively charged at neutral pH. The complete reversal of the electrostatic potential of the substrate-binding site suggests that the different Rpfs function optimally at different pHs or most efficiently hydrolyze different micro-domains of PG. These studies provide insights into the molecular determinants of the evolution of functional specialization in Rpfs.
- Department of Molecular and Cell Biology and California Institute for Quantitative Biosciences, University of California, Berkeley, California, 94720.
Organizational Affiliation: