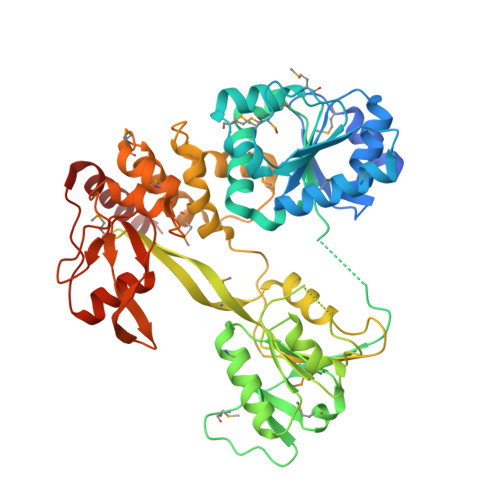
X-Ray Structure of the Pestivirus Ns3 Helicase and its Conformation in Solution.
Tortorici, M.A., Duquerroy, S., Kwok, J., Vonrhein, C., Perez, J., Lamp, B., Bricogne, G., Rumenapf, T., Vachette, P., Rey, F.A.(2015) J Virol 89: 4356
- PubMed: 25653438 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.03165-14
- Primary Citation Related Structures:
4CBG, 4CBH, 4CBI, 4CBL, 4CBM - PubMed Abstract:
Pestiviruses form a genus in the Flaviviridae family of small enveloped viruses with a positive-sense single-stranded RNA genome. Viral replication in this family requires the activity of a superfamily 2 RNA helicase contained in the C-terminal domain of nonstructural protein 3 (NS3). NS3 features two conserved RecA-like domains (D1 and D2) with ATPase activity, plus a third domain (D3) that is important for unwinding nucleic acid duplexes. We report here the X-ray structure of the pestivirus NS3 helicase domain (pNS3h) at a 2.5-Å resolution. The structure deviates significantly from that of NS3 of other genera in the Flaviviridae family in D3, as it contains two important insertions that result in a narrower nucleic acid binding groove. We also show that mutations in pNS3h that rescue viruses from which the core protein is deleted map to D3, suggesting that this domain may be involved in interactions that facilitate particle assembly. Finally, structural comparisons of the enzyme in different crystalline environments, together with the findings of small-angle X-ray-scattering studies in solution, show that D2 is mobile with respect to the rest of the enzyme, oscillating between closed and open conformations. Binding of a nonhydrolyzable ATP analog locks pNS3h in a conformation that is more compact than the closest apo-form in our crystals. Together, our results provide new insight and bring up new questions about pNS3h function during pestivirus replication. Although pestivirus infections impose an important toll on the livestock industry worldwide, little information is available about the nonstructural proteins essential for viral replication, such as the NS3 helicase. We provide here a comparative structural and functional analysis of pNS3h with respect to its orthologs in other viruses of the same family, the flaviviruses and hepatitis C virus. Our studies reveal differences in the nucleic acid binding groove that could have implications for understanding the unwinding specificity of pNS3h, which is active only on RNA duplexes. We also show that pNS3h has a highly dynamic behavior--a characteristic probably shared with NS3 helicases from all Flaviviridae members--that could be targeted for drug design by using recent algorithms to specifically block molecular motion. Compounds that lock the enzyme in a single conformation or limit its dynamic range of conformations are indeed likely to block its helicase function.
- Unité de Virologie Structurale, Département de Virologie, Institut Pasteur, Paris, France CNRS, UMR 3569, Paris, France tortoric@pasteur.fr rey@pasteur.fr.
Organizational Affiliation: