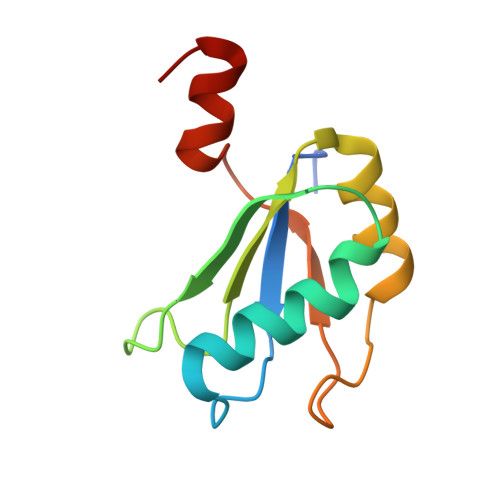
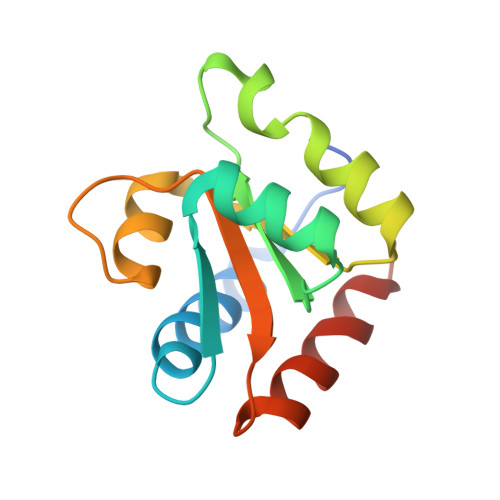
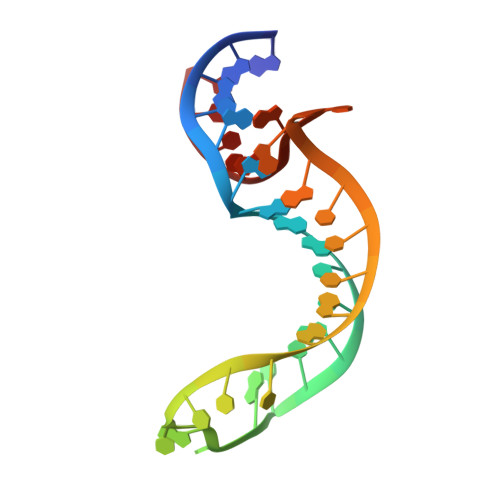
Structure of a Rare, Non-Standard Sequence K-Turn Bound by L7Ae Protein
Huang, L., Lilley, D.M.J.(2014) Nucleic Acids Res 42: 4734
- PubMed: 24482444 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku087
- Primary Citation Related Structures:
4C4W - PubMed Abstract:
Kt-23 from Thelohania solenopsae is a rare RNA kink turn (k-turn) where an adenine replaces the normal guanine at the 2n position. L7Ae is a member of a strongly conserved family of proteins that bind a range of k-turn structures in the ribosome, box C/D and H/ACA small nucleolar RNAs and U4 small nuclear RNA. We have solved the crystal structure of T. solenopsae Kt-23 RNA bound to Archeoglobus fulgidus L7Ae protein at a resolution of 2.95 Å. The protein binds in the major groove displayed on the outer face of the k-turn, in a manner similar to complexes with standard k-turn structures. The k-turn adopts a standard N3 class conformation, with a single hydrogen bond from A2b N6 to A2n N3. This contrasts with the structure of the same sequence located in the SAM-I riboswitch, where it adopts an N1 structure, showing the inherent plasticity of k-turn structure. This potentially can affect any tertiary interactions in which the RNA participates.
- Cancer Research UK Nucleic Acid Structure Research Group, MSI/WTB Complex, The University of Dundee, Dow Street, Dundee DD1 5EH, UK.
Organizational Affiliation: