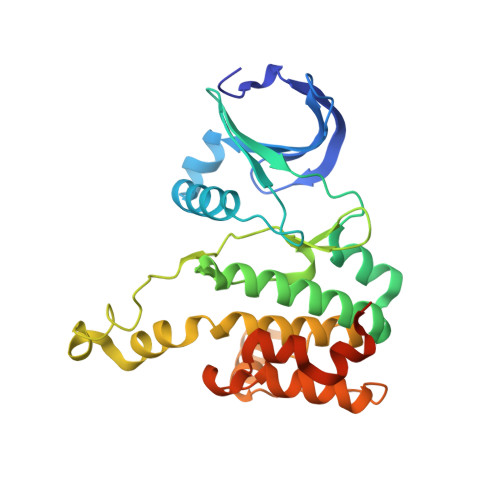
Molecular Mechanism of Aurora a Kinase Autophosphorylation and its Allosteric Activation by Tpx2.
Zorba, A., Buosi, V., Kutter, S., Kern, N., Pontiggia, F., Cho, Y.J., Kern, D.(2014) Elife 3: 02667
- PubMed: 24867643 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.02667
- Primary Citation Related Structures:
4C3P, 4C3R - PubMed Abstract:
We elucidate the molecular mechanisms of two distinct activation strategies (autophosphorylation and TPX2-mediated activation) in human Aurora A kinase. Classic allosteric activation is in play where either activation loop phosphorylation or TPX2 binding to a conserved hydrophobic groove shifts the equilibrium far towards the active conformation. We resolve the controversy about the mechanism of autophosphorylation by demonstrating intermolecular autophosphorylation in a long-lived dimer by combining X-ray crystallography with functional assays. We then address the allosteric activation by TPX2 through activity assays and the crystal structure of a domain-swapped dimer of dephosphorylated Aurora A and TPX2(1-25). While autophosphorylation is the key regulatory mechanism in the centrosomes in the early stages of mitosis, allosteric activation by TPX2 of dephosphorylated Aurora A could be at play in the spindle microtubules. The mechanistic insights into autophosphorylation and allosteric activation by TPX2 binding proposed here, may have implications for understanding regulation of other protein kinases.DOI: http://dx.doi.org/10.7554/eLife.02667.001.
- Department of Biochemistry, Howard Hughes Medical Institute, Brandeis University, Waltham, United States.
Organizational Affiliation: