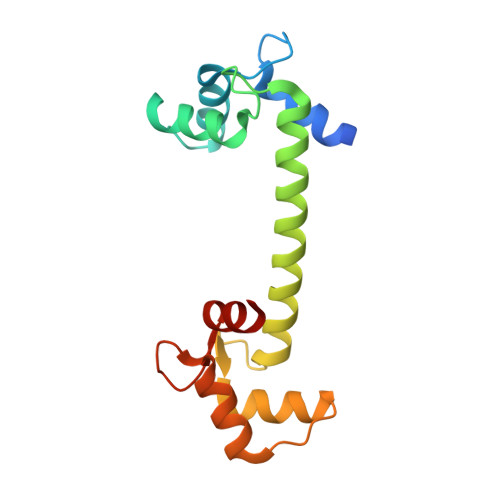
Crystallographic Snapshots of Initial Steps in the Collapse of the Calmodulin Central Helix
Kursula, P.(2014) Acta Crystallogr D Biol Crystallogr 70: 24
- PubMed: 24419375 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004713024437
- Primary Citation Related Structures:
4BW7, 4BW8 - PubMed Abstract:
Calmodulin is one of the most well characterized proteins and a widely used model system for calcium binding and large-scale protein conformational changes. Its long central helix is usually cut in half when a target peptide is bound. Here, two new crystal structures of calmodulin are presented, in which conformations possibly representing the first steps of calmodulin conformational collapse have been trapped. The central helix in the two structures is bent in the middle, causing a significant movement of the N- and C-terminal lobes with respect to one another. In both of the bent structures, a nearby polar side chain is inserted into the helical groove, disrupting backbone hydrogen bonding. The structures give an insight into the details of the factors that may be involved in the distortion of the central helix upon ligand peptide binding.
- Department of Biochemistry and Biocenter Oulu, University of Oulu, Oulu, Finland.
Organizational Affiliation: