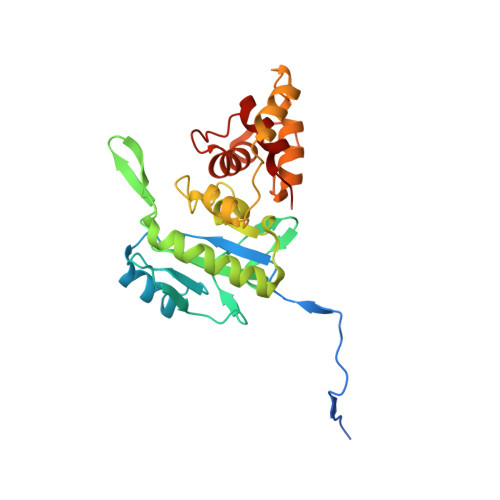
Reaction Products and the X-Ray Structure of Ampdh2, a Virulence Determinant of Pseudomonas Aeruginosa.
Martinez-Caballero, S., Lee, M., Artola-Recolons, C., Carrasco-Lopez, C., Hesek, D., Spink, E.E., Lastochkin, E., Zhang, W., Hellman, L.M., Boggess, B., Mobashery, S., Hermoso, J.A.(2013) J Am Chem Soc 135: 10318
- PubMed: 23819763 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja405464b
- Primary Citation Related Structures:
4BJ4, 4BOL, 4BPA - PubMed Abstract:
The zinc protease AmpDh2 is a virulence determinant of Pseudomonas aeruginosa, a problematic human pathogen. The mechanism of how the protease manifests virulence is not known, but it is known that it turns over the bacterial cell wall. The reaction of AmpDh2 with the cell wall was investigated, and nine distinct turnover products were characterized by LC/MS/MS. The enzyme turns over both the cross-linked and noncross-linked cell wall. Three high-resolution X-ray structures, the apo enzyme and two complexes with turnover products, were solved. The X-ray structures show how the dimeric protein interacts with the inner leaflet of the bacterial outer membrane and that the two monomers provide a more expansive surface for recognition of the cell wall. This binding surface can accommodate the 3D solution structure of the cross-linked cell wall.
- Department of Crystallography and Structural Biology, Inst. Química-Física "Rocasolano", CSIC, Serrano 119, 28006 Madrid, Spain.
Organizational Affiliation: