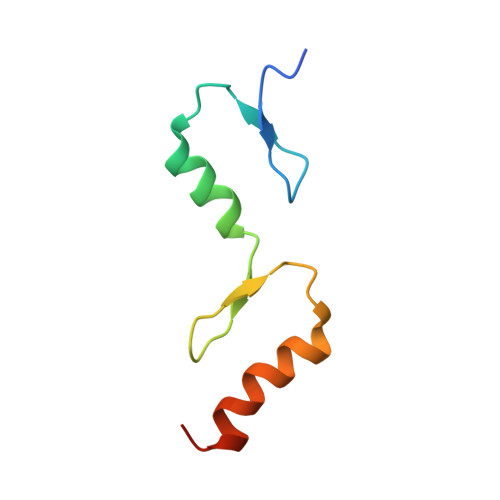
The Structure of Tax1BP1 Ubz1 + 2 Provides Insight Into Target Specificity and Adaptability
Ceregido, M.A., Spinola-Amilibia, M., Buts, L., Rivera-Torres, J., Garcia-Pino, A., Bravo, J., Van Nuland, N.A.J.(2014) J Mol Biology 426: 674
- PubMed: 24239949 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.11.006
- Primary Citation Related Structures:
2M7Q, 4BMJ - PubMed Abstract:
TAX1BP1 is a novel ubiquitin-binding adaptor protein involved in the negative regulation of the NF-kappaB transcription factor, which is a key player in inflammatory responses, immunity and tumorigenesis. TAX1BP1 recruits A20 to the ubiquitinated signaling proteins TRAF6 and RIP1, leading to their A20-mediated deubiquitination and the disruption of IL-1-induced and TNF-induced NF-kappaB signaling, respectively. The two zinc fingers localized at its C-terminus function as novel ubiquitin-binding domains (UBZ, ubiquitin-binding zinc finger). Here we present for the first time both the solution and crystal structures of two classical UBZ domains in tandem within the human TAX1BP1. The relative orientation of the two domains is slightly different in the X-ray structure with respect to the NMR structure, indicating some degree of conformational flexibility, which is rationalized by NMR relaxation data. The observed degree of flexibility and stability between the two UBZ domains might have consequences on the recognition mechanism of interacting partners.
- Departamento de Química Física e Instituto de Biotecnología, Facultad de Ciencias, Universidad de Granada, Granada 18071, Spain; Jean Jeener NMR Centre, Structural Biology Brussels, Vrije Universiteit Brussel, Brussels 1050, Belgium; Molecular Recognition Unit, Department of Structural Biology, VIB, Brussels 1050, Belgium.
Organizational Affiliation: