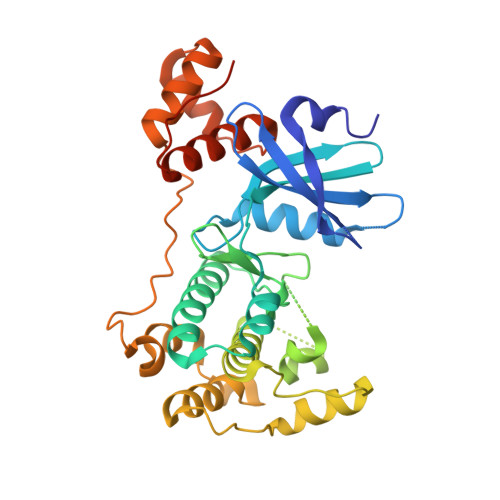
Structural Insight Into Maternal Embryonic Leucine Zipper Kinase (Melk) Conformation and Inhibition Towards Structure- Based Drug Design.
Canevari, G., Re Depaolini, S., Cucchi, U., Bertrand, J.A., Casale, E., Perrera, C., Forte, B., Carpinelli, P., Felder, E.R.(2013) Biochemistry 52: 6380
- PubMed: 23914841 Search on PubMed
- DOI: https://doi.org/10.1021/bi4005864
- Primary Citation Related Structures:
4BKY, 4BKZ - PubMed Abstract:
Maternal embryonic leucine zipper kinase (MELK) is upregulated in several types of tumor, including breast, prostate, and brain tumors. Its expression is generally associated with cell survival, cell proliferation, and resistance to apoptosis. Therefore, the potential of MELK inhibitors as therapeutic agents is recently attracting considerable interest. Here we report the first structures of MELK in complex with AMP-PNP and with nanomolar inhibitors. Our studies shed light on the role of the MELK UBA domain, provide a characterization of the kinase active site, and identify key residues for achieving high potency, laying the groundwork for structure-based drug design efforts.
- Nerviano Medical Sciences , Viale Pasteur 10, 20014 Nerviano, Milan, Italy.
Organizational Affiliation: