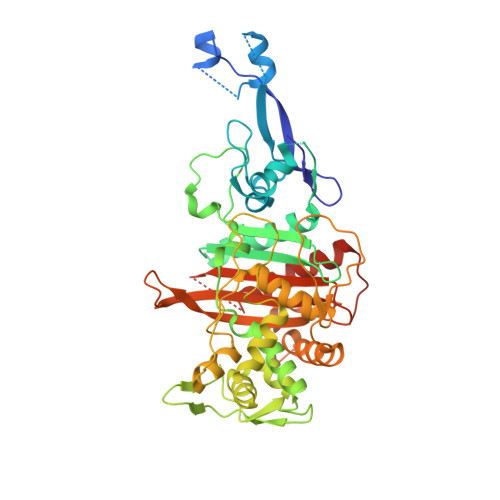
Crystal Structure of Penicillin-Binding Protein 3 (Pbp3) from Escherichia Coli.
Sauvage, E., Derouaux, A., Fraipont, C., Joris, M., Herman, R., Rocaboy, M., Schloesser, M., Dumas, J., Kerff, F., Nguyen-Disteche, M., Charlier, P.(2014) PLoS One 9: 98042
- PubMed: 24875494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0098042
- Primary Citation Related Structures:
4BJP, 4BJQ - PubMed Abstract:
In Escherichia coli, penicillin-binding protein 3 (PBP3), also known as FtsI, is a central component of the divisome, catalyzing cross-linking of the cell wall peptidoglycan during cell division. PBP3 is mainly periplasmic, with a 23 residues cytoplasmic tail and a single transmembrane helix. We have solved the crystal structure of a soluble form of PBP3 (PBP3(57-577)) at 2.5 Å revealing the two modules of high molecular weight class B PBPs, a carboxy terminal module exhibiting transpeptidase activity and an amino terminal module of unknown function. To gain additional insight, the PBP3 Val88-Ser165 subdomain (PBP3(88-165)), for which the electron density is poorly defined in the PBP3 crystal, was produced and its structure solved by SAD phasing at 2.1 Å. The structure shows a three dimensional domain swapping with a β-strand of one molecule inserted between two strands of the paired molecule, suggesting a possible role in PBP3(57-577) dimerization.
- Centre d'Ingénierie des Protéines, Université de Liège, Institut de Physique B5a et Institut de Chimie B6a, Sart Tilman, Liège, Belgium.
Organizational Affiliation: