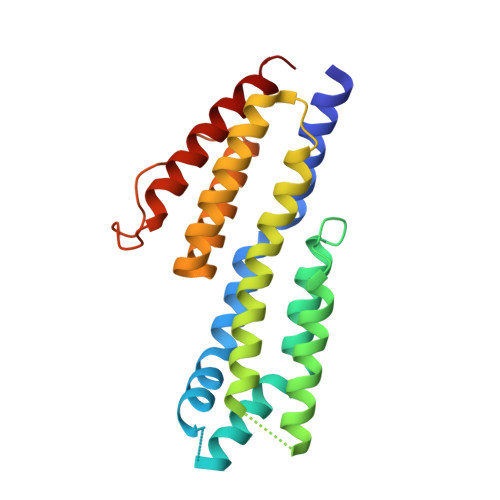
Structural Characterization of Cspz, a Complement Regulator Factor H and Fhl-1 Binding Protein from Borrelia Burgdorferi.
Brangulis, K., Petrovskis, I., Kazaks, A., Bogans, J., Otikovs, M., Jaudzems, K., Ranka, R., Tars, K.(2014) FEBS J 281: 2613
- PubMed: 24702793 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12808
- Primary Citation Related Structures:
4BG0, 4CBE - PubMed Abstract:
Borrelia burgdorferi is the causative agent of Lyme disease and is found in two different types of hosts in nature - Ixodes ticks and various mammalian organisms. To initiate disease and survive in mammalian host organisms, B. burgdorferi must be able to transfer to a new host, proliferate, attach to different tissue and resist the immune response. To resist the host's immune response, B. burgdorferi produces at least five different outer surface proteins that can bind complement regulator factor H (CFH) and/or factor H-like protein 1 (CFHL-1). The crystal structures of two uniquely folded complement binding proteins, which belong to two distinct gene families and are not found in other bacteria, have been previously described. The crystal structure of the CFH and CFHL-1 binding protein CspZ (also known as BbCRASP-2 or BBH06) from B. burgdorferi, which belongs to a third gene family, is reported in this study. The structure reveals that the overall fold is different from the known structures of the other complement binding proteins in B. burgdorferi or other bacteria; this structure does not resemble the fold of any known protein deposited in the Protein Data Bank. The N-terminal part of the CspZ protein forms a four-helix bundle and has features similar to the FAT domain (focal adhesion targeting domain) and a related domain found in the vinculin/α-catenin family. By combining our findings from the crystal structure of CspZ with previous mutagenesis studies, we have identified a likely binding surface on CspZ for CFH and CFHL-1.
- Latvian Biomedical Research and Study Centre, Riga, Latvia; Riga Stradins University, Latvia.
Organizational Affiliation: