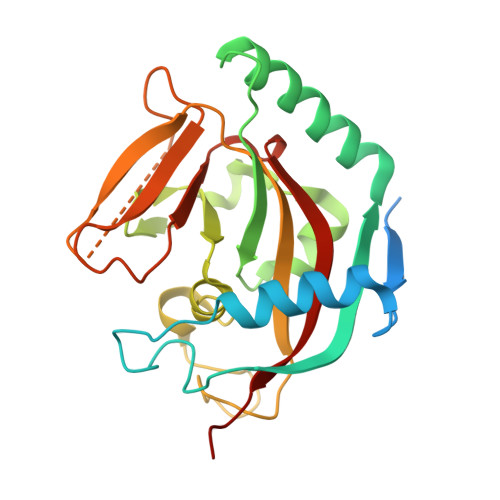
Structural Basis and Selectivity of Tankyrase Inhibition by a Wnt Signaling Inhibitor Wiki4
Haikarainen, T., Venkannagari, H., Narwal, M., Obaji, E., Lee, H., Nkizinkiko, Y., Lehtio, L.(2013) PLoS One 8: 65404
- PubMed: 23762361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0065404
- Primary Citation Related Structures:
4BFP - PubMed Abstract:
Recently a novel inhibitor of Wnt signaling was discovered. The compound, WIKI4, was found to act through tankyrase inhibition and regulate β-catenin levels in many cancer cell lines and human embryonic stem cells. Here we confirm that WIKI4 is a high potency tankyrase inhibitor and that it selectively inhibits tankyrases over other ARTD enzymes tested. The binding mode of the compound to tankyrase 2 was determined by protein X-ray crystallography to 2.4 Å resolution. The structure revealed a novel binding mode to the adenosine subsite of the donor NAD(+) binding groove of the catalytic domain. Our results form a structural basis for further development of potent and selective tankyrase inhibitors based on the WIKI4 scaffold.
- Biocenter Oulu, Department of Biochemistry, University of Oulu, Oulu, Finland.
Organizational Affiliation: