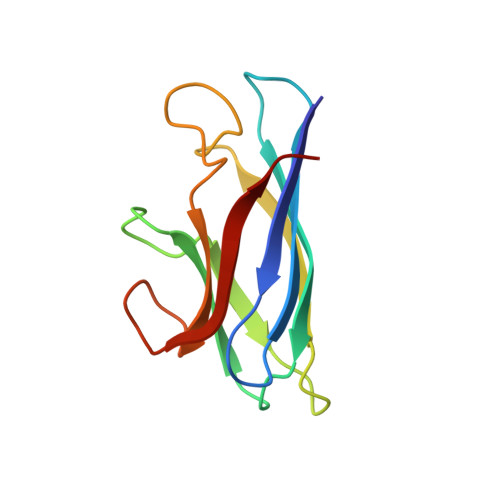
Global Structural Motions from the Strain of a Single Hydrogen Bond.
Danielsson, J., Awad, W., Saraboji, K., Kurnik, M., Lang, L., Leinartaite, L., Marklund, S.L., Logan, D.T., Oliveberg, M.(2013) Proc Natl Acad Sci U S A 110: 3829
- PubMed: 23431167 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1217306110
- Primary Citation Related Structures:
4BCY, 4BCZ, 4BD4 - PubMed Abstract:
The origin and biological role of dynamic motions of folded enzymes is not yet fully understood. In this study, we examine the molecular determinants for the dynamic motions within the β-barrel of superoxide dismutase 1 (SOD1), which previously were implicated in allosteric regulation of protein maturation and also pathological misfolding in the neurodegenerative disease amyotrophic lateral sclerosis. Relaxation-dispersion NMR, hydrogen/deuterium exchange, and crystallographic data show that the dynamic motions are induced by the buried H43 side chain, which connects the backbones of the Cu ligand H120 and T39 by a hydrogen-bond linkage through the hydrophobic core. The functional role of this highly conserved H120-H43-T39 linkage is to strain H120 into the correct geometry for Cu binding. Upon elimination of the strain by mutation H43F, the apo protein relaxes through hydrogen-bond swapping into a more stable structure and the dynamic motions freeze out completely. At the same time, the holo protein becomes energetically penalized because the twisting back of H120 into Cu-bound geometry leads to burial of an unmatched backbone carbonyl group. The question then is whether this coupling between metal binding and global structural motions in the SOD1 molecule is an adverse side effect of evolving viable Cu coordination or plays a key role in allosteric regulation of biological function, or both?
- Department of Biochemistry and Biophysics, Arrhenius Laboratories of Natural Sciences, Stockholm University, S-106 91 Stockholm, Sweden. jensd@dbb.su.se
Organizational Affiliation: